Automated breast mass detection in 3D reconstructed tomosynthesis volumes: a featureless approach
- PMID: 18777923
- PMCID: PMC2673649
- DOI: 10.1118/1.2953562
Automated breast mass detection in 3D reconstructed tomosynthesis volumes: a featureless approach
Abstract
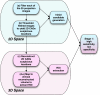
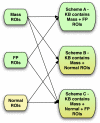
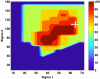
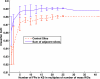
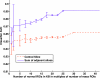
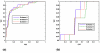
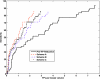
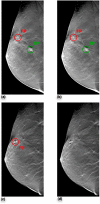
The purpose of this study was to propose and implement a computer aided detection (CADe) tool for breast tomosynthesis. This task was accomplished in two stages-a highly sensitive mass detector followed by a false positive (FP) reduction stage. Breast tomosynthesis data from 100 human subject cases were used, of which 25 subjects had one or more mass lesions and the rest were normal. For stage 1, filter parameters were optimized via a grid search. The CADe identified suspicious locations were reconstructed to yield 3D CADe volumes of interest. The first stage yielded a maximum sensitivity of 93% with 7.7 FPs/breast volume. Unlike traditional CADe algorithms in which the second stage FP reduction is done via feature extraction and analysis, instead information theory principles were used with mutual information as a similarity metric. Three schemes were proposed, all using leave-one-case-out cross validation sampling. The three schemes, A, B, and C, differed in the composition of their knowledge base of regions of interest (ROIs). Scheme A's knowledge base was comprised of all the mass and FP ROIs generated by the first stage of the algorithm. Scheme B had a knowledge base that contained information from mass ROIs and randomly extracted normal ROIs. Scheme C had information from three sources of information-masses, FPs, and normal ROIs. Also, performance was assessed as a function of the composition of the knowledge base in terms of the number of FP or normal ROIs needed by the system to reach optimal performance. The results indicated that the knowledge base needed no more than 20 times as many FPs and 30 times as many normal ROIs as masses to attain maximal performance. The best overall system performance was 85% sensitivity with 2.4 FPs per breast volume for scheme A, 3.6 FPs per breast volume for scheme B, and 3 FPs per breast volume for scheme C.
Figures

Similar articles
-
Computer-aided detection of clustered microcalcifications in digital breast tomosynthesis: a 3D approach.Med Phys. 2012 Jan;39(1):28-39. doi: 10.1118/1.3662072. Med Phys. 2012. PMID: 22225272 Free PMC article.
-
Multi-domain features for reducing false positives in automated detection of clustered microcalcifications in digital breast tomosynthesis.Med Phys. 2019 Mar;46(3):1300-1308. doi: 10.1002/mp.13394. Epub 2019 Feb 14. Med Phys. 2019. PMID: 30661242
-
Computer aided detection of clusters of microcalcifications on full field digital mammograms.Med Phys. 2006 Aug;33(8):2975-88. doi: 10.1118/1.2211710. Med Phys. 2006. PMID: 16964876
-
Local Binary Patterns Descriptor Based on Sparse Curvelet Coefficients for False-Positive Reduction in Mammograms.J Healthc Eng. 2018 Sep 25;2018:5940436. doi: 10.1155/2018/5940436. eCollection 2018. J Healthc Eng. 2018. PMID: 30356422 Free PMC article.
-
CADx of mammographic masses and clustered microcalcifications: a review.Med Phys. 2009 Jun;36(6):2052-68. doi: 10.1118/1.3121511. Med Phys. 2009. PMID: 19610294 Review.
Cited by
-
Digital breast tomosynthesis: lessons learned from early clinical implementation.Radiographics. 2014 Jul-Aug;34(4):E89-102. doi: 10.1148/rg.344130087. Radiographics. 2014. PMID: 25019451 Free PMC article.
-
Computer-aided detection of clustered microcalcifications in digital breast tomosynthesis: a 3D approach.Med Phys. 2012 Jan;39(1):28-39. doi: 10.1118/1.3662072. Med Phys. 2012. PMID: 22225272 Free PMC article.
-
Clinical implementation of digital breast tomosynthesis.Radiol Clin North Am. 2014 May;52(3):499-518. doi: 10.1016/j.rcl.2013.11.013. Epub 2014 Feb 18. Radiol Clin North Am. 2014. PMID: 24792652 Free PMC article. Review.
-
A review of breast tomosynthesis. Part II. Image reconstruction, processing and analysis, and advanced applications.Med Phys. 2013 Jan;40(1):014302. doi: 10.1118/1.4770281. Med Phys. 2013. PMID: 23298127 Free PMC article. Review.
-
Automated detection of mass lesions in dedicated breast CT: a preliminary study.Med Phys. 2012 Feb;39(2):866-73. doi: 10.1118/1.3678991. Med Phys. 2012. PMID: 22320796 Free PMC article.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

