An unusual intrinsically disordered protein from the model legume Lotus japonicus stabilizes proteins in vitro
- PMID: 18779323
- PMCID: PMC2662180
- DOI: 10.1074/jbc.M805024200
An unusual intrinsically disordered protein from the model legume Lotus japonicus stabilizes proteins in vitro
Abstract
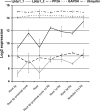
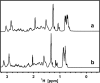
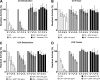
Intrinsic structural disorder is a prevalent feature of proteins with chaperone activity. Using a complementary set of techniques, we have structurally characterized LjIDP1 (intrinsically disordered protein 1) from the model legume Lotus japonicus, and our results provide the first structural characterization of a member of the Lea5 protein family (PF03242). Contrary to in silico predictions, we show that LjIDP1 is intrinsically disordered and probably exists as an ensemble of conformations with limited residual beta-sheet, turn/loop, and polyproline II secondary structure. Furthermore, we show that LjIDP1 has an inherent propensity to undergo a large conformational shift, adopting a largely alpha-helical structure when it is dehydrated and in the presence of different detergents and alcohols. This is consistent with an overrepresentation of order-promoting residues in LjIDP1 compared with the average of intrinsically disordered proteins. In line with functioning as a chaperone, we show that LjIDP1 effectively prevents inactivation of two model enzymes under conditions that promote protein misfolding and aggregation. The LjIdp1 gene is expressed in all L. japonicus tissues tested. A higher expression level was found in the root tip proximal zone, in roots inoculated with compatible endosymbiotic M. loti, and in functional nitrogen-fixing root nodules. We suggest that the ability of LjIDP1 to prevent protein misfolding and aggregation may play a significant role in tissues, such as symbiotic root nodules, which are characterized by high metabolic activity.
Figures





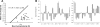

References
-
- Uversky, V. N., Oldfield, C. J., and Dunker, A. K. (2008) Annu. Rev. Biophys. 37 215-246 - PubMed
-
- Tunnacliffe, A., and Wise, M. J. (2007) Naturwissenschaften 94 791-812 - PubMed
-
- Dyson, H. J., and Wright, P. E. (2005) Nat. Rev. Mol. Cell Biol. 6 197-208 - PubMed
-
- Colebatch, G., Desbrosses, G., Ott, T., Krusell, L., Montanari, O., Kloska, S., Kopka, J., and Udvardi, M. K. (2004) Plant J. 39 487-512 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

