End joining at Caenorhabditis elegans telomeres
- PMID: 18780750
- PMCID: PMC2567377
- DOI: 10.1534/genetics.108.089920
End joining at Caenorhabditis elegans telomeres
Abstract
Critically shortened telomeres can be subjected to DNA repair events that generate end-to-end chromosome fusions. The resulting dicentric chromosomes can enter breakage-fusion-bridge cycles, thereby impeding elucidation of the structures of the initial fusion events and a mechanistic understanding of their genesis. Current models for the molecular basis of fusion of critically shortened, uncapped telomeres rely on PCR assays that typically capture fusion breakpoints created by direct ligation of chromosome ends. Here we use independent approaches that rely on distinctive features of Caenorhabditis elegans to study the frequency of direct end-to-end chromosome fusion in telomerase mutants: (1) holocentric chromosomes that allow for genetic isolation of stable end-to-end fusions and (2) unique subtelomeric sequences that allow for thorough PCR analysis of samples of genomic DNA harboring multiple end-to-end fusions. Surprisingly, only a minority of end-to-end fusion events resulted from direct end joining with no additional genome rearrangements. We also demonstrate that deficiency for the C. elegans Ku DNA repair heterodimer does not affect telomere length or cause synthetic effects in the absence of telomerase.
Figures




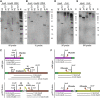
Similar articles
-
DNA synthesis generates terminal duplications that seal end-to-end chromosome fusions.Science. 2011 Apr 22;332(6028):468-71. doi: 10.1126/science.1199022. Science. 2011. PMID: 21512032 Free PMC article.
-
Spontaneous telomere to telomere fusions occur in unperturbed fission yeast cells.Nucleic Acids Res. 2013 Mar 1;41(5):3056-67. doi: 10.1093/nar/gks1459. Epub 2013 Jan 18. Nucleic Acids Res. 2013. PMID: 23335786 Free PMC article.
-
Caenorhabditis elegans POT-1 and POT-2 repress telomere maintenance pathways.G3 (Bethesda). 2013 Feb;3(2):305-13. doi: 10.1534/g3.112.004440. Epub 2013 Feb 1. G3 (Bethesda). 2013. PMID: 23390606 Free PMC article.
-
Repair and Reconstruction of Telomeric and Subtelomeric Regions and Genesis of New Telomeres: Implications for Chromosome Evolution.Bioessays. 2020 Jun;42(6):e1900177. doi: 10.1002/bies.201900177. Epub 2020 Apr 1. Bioessays. 2020. PMID: 32236965 Review.
-
Telomeres and telomerase.Philos Trans R Soc Lond B Biol Sci. 2004 Jan 29;359(1441):109-21. doi: 10.1098/rstb.2003.1370. Philos Trans R Soc Lond B Biol Sci. 2004. PMID: 15065663 Free PMC article. Review.
Cited by
-
Long-read sequencing and de novo genome assemblies reveal complex chromosome end structures caused by telomere dysfunction at the single nucleotide level.Nucleic Acids Res. 2021 Apr 6;49(6):3338-3353. doi: 10.1093/nar/gkab141. Nucleic Acids Res. 2021. PMID: 33693840 Free PMC article.
-
Telomere disruption results in non-random formation of de novo dicentric chromosomes involving acrocentric human chromosomes.PLoS Genet. 2010 Aug 12;6(8):e1001061. doi: 10.1371/journal.pgen.1001061. PLoS Genet. 2010. PMID: 20711355 Free PMC article.
-
Binding of an X-Specific Condensin Correlates with a Reduction in Active Histone Modifications at Gene Regulatory Elements.Genetics. 2019 Jul;212(3):729-742. doi: 10.1534/genetics.119.302254. Epub 2019 May 22. Genetics. 2019. PMID: 31123040 Free PMC article.
-
Heterologous synapsis in C. elegans is regulated by meiotic double-strand breaks and crossovers.Chromosoma. 2021 Dec;130(4):237-250. doi: 10.1007/s00412-021-00763-y. Epub 2021 Oct 4. Chromosoma. 2021. PMID: 34608541 Free PMC article.
-
Synthetic cytotoxicity: digenic interactions with TEL1/ATM mutations reveal sensitivity to low doses of camptothecin.Genetics. 2014 Jun;197(2):611-23. doi: 10.1534/genetics.114.161307. Epub 2014 Mar 20. Genetics. 2014. PMID: 24653001 Free PMC article.
References
-
- Ahmed, S., A. Alpi, M. O. Hengartner and A. Gartner, 2001. C. elegans RAD-5/CLK-2 defines a new DNA damage checkpoint protein. Curr. Biol. 11 1934–1944. - PubMed
-
- Ahmed, S., and J. Hodgkin, 2000. MRT-2 checkpoint protein is required for germline immortality and telomere replication in C. elegans. Nature 403 159–164. - PubMed
-
- Bae, N. S., and P. Baumann, 2007. A RAP1/TRF2 complex inhibits nonhomologous end-joining at human telomeric DNA ends. Mol. Cell 26 323–334. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

