Selection of internal reference genes for SYBR green qRT-PCR studies of rhesus monkey (Macaca mulatta) tissues
- PMID: 18782457
- PMCID: PMC2561044
- DOI: 10.1186/1471-2199-9-78
Selection of internal reference genes for SYBR green qRT-PCR studies of rhesus monkey (Macaca mulatta) tissues
Abstract
Background: The rhesus monkey (Macaca mulatta) is a valuable and widely used model animal for biomedical research. However, quantitative analyses of rhesus gene expression profiles under diverse experimental conditions are limited by a shortage of suitable internal controls for the normalization of mRNA levels. In this study, we used a systematic approach for the selection of potential reference genes in the rhesus monkey and compared their suitability to that of the corresponding genes in humans.
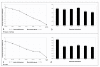
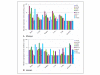
Results: Eight housekeeping genes (HKGs) (GAPDH, SDHA, ACTB, RPL13A, RPL32, UBA52, PGK1Y, and YWHAZ) from rhesus monkeys and humans were selected to test for normalization of expression levels in six different tissue types (brain, colon, kidney, liver, lung, and stomach). Their stability and suitability as reference genes were validated by geNorm, NormFinder and BestKeeper programs. Intriguingly, RPL13A and RPL32 were selected as ideal reference genes only in rhesus monkeys.
Conclusion: The results clearly indicated the necessity of using different reference genes for normalization of expression levels between rhesus monkeys and humans in various tissues.
Figures
Similar articles
-
Microarray analysis of relative gene expression stability for selection of internal reference genes in the rhesus macaque brain.BMC Mol Biol. 2010 Jun 21;11:47. doi: 10.1186/1471-2199-11-47. BMC Mol Biol. 2010. PMID: 20565976 Free PMC article.
-
PSMB2 and RPL32 are suitable denominators to normalize gene expression profiles in bronchoalveolar cells.BMC Mol Biol. 2008 Jul 31;9:69. doi: 10.1186/1471-2199-9-69. BMC Mol Biol. 2008. PMID: 18671841 Free PMC article.
-
Selection of appropriate reference genes for RT-qPCR analysis in a streptozotocin-induced Alzheimer's disease model of cynomolgus monkeys (Macaca fascicularis).PLoS One. 2013;8(2):e56034. doi: 10.1371/journal.pone.0056034. Epub 2013 Feb 14. PLoS One. 2013. PMID: 23457495 Free PMC article.
-
Evaluation of reference genes for human chondrocytes cultured in several different thermal environments.Int J Hyperthermia. 2014 May;30(3):210-6. doi: 10.3109/02656736.2014.906048. Int J Hyperthermia. 2014. PMID: 24773042
-
Advancing primatology through ethical and scientific perspectives on rhesus monkey (Macaca mulatta) cloning.J Med Primatol. 2024 Jun;53(3):e12704. doi: 10.1111/jmp.12704. J Med Primatol. 2024. PMID: 38812105 Review.
Cited by
-
Exogenous reference gene normalization for real-time reverse transcription-polymerase chain reaction analysis under dynamic endogenous transcription.Neural Regen Res. 2012 May 15;7(14):1064-72. doi: 10.3969/j.issn.1673-5374.2012.14.004. Neural Regen Res. 2012. PMID: 25722696 Free PMC article.
-
Genome-wide analysis of DNA methylation in human amnion.ScientificWorldJournal. 2013;2013:678156. doi: 10.1155/2013/678156. Epub 2013 Feb 19. ScientificWorldJournal. 2013. PMID: 23533356 Free PMC article.
-
Behavioral inhibition in rhesus monkeys (Macaca mulatta) is related to the airways response, but not immune measures, commonly associated with asthma.PLoS One. 2013 Aug 9;8(8):e71575. doi: 10.1371/journal.pone.0071575. eCollection 2013. PLoS One. 2013. PMID: 23951195 Free PMC article.
-
Assessment of Aedes albopictus reference genes for quantitative PCR at different stages of development.PLoS One. 2018 Mar 19;13(3):e0194664. doi: 10.1371/journal.pone.0194664. eCollection 2018. PLoS One. 2018. PMID: 29554153 Free PMC article.
-
Serum levels of microRNAs can specifically predict liver injury of chronic hepatitis B.World J Gastroenterol. 2012 Oct 7;18(37):5188-96. doi: 10.3748/wjg.v18.i37.5188. World J Gastroenterol. 2012. PMID: 23066312 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

