De novo design and evolution of artificial disulfide isomerase enzymes analogous to the bacterial DsbC
- PMID: 18782764
- PMCID: PMC2581571
- DOI: 10.1074/jbc.M803346200
De novo design and evolution of artificial disulfide isomerase enzymes analogous to the bacterial DsbC
Abstract
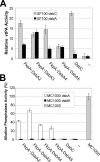
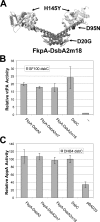
The Escherichia coli disulfide isomerase, DsbC is a V-shaped homodimer with each monomer comprising a dimerization region that forms part of a putative peptide-binding pocket and a thioredoxin catalytic domain. Disulfide isomerases from prokaryotes and eukaryotes exhibit little sequence homology but display very similar structural organization with two thioredoxin domains facing each other on top of the dimerization/peptide-binding region. To aid the understanding of the mechanistic significance of thioredoxin domain dimerization and of the peptide-binding cleft of DsbC, we constructed a series of protein chimeras comprising unrelated protein dimerization domains fused to thioredoxin superfamily enzymes. Chimeras consisting of the dimerization domain and the alpha-helical linker of the bacterial proline cis/trans isomerase FkpA and the periplasmic oxidase DsbA gave rise to enzymes that catalyzed the folding of multidisulfide substrate proteins in vivo with comparable efficiency to E. coli DsbC. In addition, expression of FkpA-DsbAs conferred modest resistance to CuCl2, a phenotype that depends on disulfide bond isomerization. Selection for resistance to elevated CuCl2 concentrations led to the isolation of FkpA-DsbA mutants containing a single amino acid substitution that changed the active site of the DsbA domain from CPHC into CPYC, increasing the similarity to the DsbC active site (CGYC). Unlike DsbC, which is resistant to oxidation by DsbB-DsbA and does not normally catalyze disulfide bond formation under physiological conditions, the FkpA-DsbA chimeras functioned both as oxidases and isomerases. The engineering of these efficient artificial isomerases delineates the key features of catalysis of disulfide bond isomerization and enhances our understanding of its evolution.
Figures

Similar articles
-
Role of dimerization in the catalytic properties of the Escherichia coli disulfide isomerase DsbC.J Biol Chem. 2009 Sep 4;284(36):23972-9. doi: 10.1074/jbc.M109.010199. Epub 2009 Jul 6. J Biol Chem. 2009. PMID: 19581640 Free PMC article.
-
Engineered DsbC chimeras catalyze both protein oxidation and disulfide-bond isomerization in Escherichia coli: Reconciling two competing pathways.Proc Natl Acad Sci U S A. 2004 Jul 6;101(27):10018-23. doi: 10.1073/pnas.0403003101. Epub 2004 Jun 25. Proc Natl Acad Sci U S A. 2004. PMID: 15220477 Free PMC article.
-
Mutants in DsbB that appear to redirect oxidation through the disulfide isomerization pathway.J Mol Biol. 2008 Apr 11;377(5):1433-42. doi: 10.1016/j.jmb.2008.01.058. Epub 2008 Jan 31. J Mol Biol. 2008. PMID: 18325532 Free PMC article.
-
Protein Disulfide Exchange by the Intramembrane Enzymes DsbB, DsbD, and CcdA.J Mol Biol. 2020 Aug 21;432(18):5091-5103. doi: 10.1016/j.jmb.2020.04.008. Epub 2020 Apr 16. J Mol Biol. 2020. PMID: 32305461 Free PMC article. Review.
-
[Escherichia coli disulfide-forming related proteins: structures, functions and their application in gene engineering for expressing heterologous proteins in Escherichia coli].Sheng Wu Gong Cheng Xue Bao. 2002 May;18(3):261-6. Sheng Wu Gong Cheng Xue Bao. 2002. PMID: 12192853 Review. Chinese.
Cited by
-
FipB, an essential virulence factor of Francisella tularensis subsp. tularensis, has dual roles in disulfide bond formation.J Bacteriol. 2014 Oct;196(20):3571-81. doi: 10.1128/JB.01359-13. Epub 2014 Aug 4. J Bacteriol. 2014. PMID: 25092026 Free PMC article.
-
C8J_1298, a bifunctional thiol oxidoreductase of Campylobacter jejuni, affects Dsb (disulfide bond) network functioning.PLoS One. 2020 Mar 23;15(3):e0230366. doi: 10.1371/journal.pone.0230366. eCollection 2020. PLoS One. 2020. PMID: 32203539 Free PMC article.
-
Strategies for successful recombinant expression of disulfide bond-dependent proteins in Escherichia coli.Microb Cell Fact. 2009 May 14;8:26. doi: 10.1186/1475-2859-8-26. Microb Cell Fact. 2009. PMID: 19442264 Free PMC article.
-
Thermal-induced dissociation and unfolding of homodimeric DsbC revealed by temperature-jump time-resolved infrared spectra.Biophys J. 2009 Nov 18;97(10):2811-9. doi: 10.1016/j.bpj.2009.08.049. Biophys J. 2009. PMID: 19917236 Free PMC article.
-
Role of dimerization in the catalytic properties of the Escherichia coli disulfide isomerase DsbC.J Biol Chem. 2009 Sep 4;284(36):23972-9. doi: 10.1074/jbc.M109.010199. Epub 2009 Jul 6. J Biol Chem. 2009. PMID: 19581640 Free PMC article.
References
-
- Kadokura, H., Katzen, F., and Beckwith, J. (2003) Annu. Rev. Biochem. 72 111–135 - PubMed
-
- Collet, J. F., and Bardwell, J. C. (2002) Mol. Microbiol. 44 1–8 - PubMed
-
- Berkmen, M., Boyd, D., and Beckwith, J. (2005) J. Biol. Chem. 280 11387–11394 - PubMed
-
- Messens, J., and Collet, J. F. (2006) Int. J. Biochem. Cell Biol. 38 1050–1062 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

