Conserved chromosomal clustering of genes governed by chromatin regulators in Drosophila
- PMID: 18783608
- PMCID: PMC2592712
- DOI: 10.1186/gb-2008-9-9-r134
Conserved chromosomal clustering of genes governed by chromatin regulators in Drosophila
Abstract
Background: The trithorax group (trxG) and Polycomb group (PcG) proteins are responsible for the maintenance of stable transcriptional patterns of many developmental regulators. They bind to specific regions of DNA and direct the post-translational modifications of histones, playing a role in the dynamics of chromatin structure.
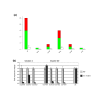
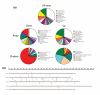
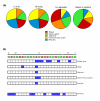
Results: We have performed genome-wide expression studies of trx and ash2 mutants in Drosophila melanogaster. Using computational analysis of our microarray data, we have identified 25 clusters of genes potentially regulated by TRX. Most of these clusters consist of genes that encode structural proteins involved in cuticle formation. This organization appears to be a distinctive feature of the regulatory networks of TRX and other chromatin regulators, since we have observed the same arrangement in clusters after experiments performed with ASH2, as well as in experiments performed by others with NURF, dMyc, and ASH1. We have also found many of these clusters to be significantly conserved in D. simulans, D. yakuba, D. pseudoobscura and partially in Anopheles gambiae.
Conclusion: The analysis of genes governed by chromatin regulators has led to the identification of clusters of functionally related genes conserved in other insect species, suggesting this chromosomal organization is biologically important. Moreover, our results indicate that TRX and other chromatin regulators may act globally on chromatin domains that contain transcriptionally co-regulated genes.
Figures






Similar articles
-
Drosophila Kismet regulates histone H3 lysine 27 methylation and early elongation by RNA polymerase II.PLoS Genet. 2008 Oct;4(10):e1000217. doi: 10.1371/journal.pgen.1000217. Epub 2008 Oct 10. PLoS Genet. 2008. PMID: 18846226 Free PMC article.
-
Genome-wide RNAi screen in Drosophila reveals Enok as a novel trithorax group regulator.Epigenetics Chromatin. 2019 Sep 23;12(1):55. doi: 10.1186/s13072-019-0301-x. Epigenetics Chromatin. 2019. PMID: 31547845 Free PMC article.
-
Dynamic Competition of Polycomb and Trithorax in Transcriptional Programming.Annu Rev Biochem. 2020 Jun 20;89:235-253. doi: 10.1146/annurev-biochem-120219-103641. Epub 2020 Jan 13. Annu Rev Biochem. 2020. PMID: 31928411 Free PMC article. Review.
-
Alternative splicing of NURF301 generates distinct NURF chromatin remodeling complexes with altered modified histone binding specificities.PLoS Genet. 2009 Jul;5(7):e1000574. doi: 10.1371/journal.pgen.1000574. Epub 2009 Jul 24. PLoS Genet. 2009. PMID: 19629165 Free PMC article.
-
Polycomb and Trithorax Group Genes in Drosophila.Genetics. 2017 Aug;206(4):1699-1725. doi: 10.1534/genetics.115.185116. Genetics. 2017. PMID: 28778878 Free PMC article. Review.
Cited by
-
Developmental Coordination of Gene Expression between Synaptic Partners During GABAergic Circuit Assembly in Cerebellar Cortex.Front Neural Circuits. 2012 Jun 26;6:37. doi: 10.3389/fncir.2012.00037. eCollection 2012. Front Neural Circuits. 2012. PMID: 22754500 Free PMC article.
-
Three Dimensional Organization of Genome Might Have Guided the Dynamics of Gene Order Evolution in Eukaryotes.Genome Biol Evol. 2016 Apr 6;8(3):946-54. doi: 10.1093/gbe/evw050. Genome Biol Evol. 2016. PMID: 26957031 Free PMC article.
-
Genomic Clustering of differential DNA methylated regions (epimutations) associated with the epigenetic transgenerational inheritance of disease and phenotypic variation.BMC Genomics. 2016 Jun 1;17:418. doi: 10.1186/s12864-016-2748-5. BMC Genomics. 2016. PMID: 27245821 Free PMC article.
-
Support for multiple classes of local expression clusters in Drosophila melanogaster, but no evidence for gene order conservation.Genome Biol. 2011;12(3):R23. doi: 10.1186/gb-2011-12-3-r23. Epub 2011 Mar 17. Genome Biol. 2011. PMID: 21414197 Free PMC article.
-
Arm-specific dynamics of chromosome evolution in malaria mosquitoes.BMC Evol Biol. 2011 Apr 7;11:91. doi: 10.1186/1471-2148-11-91. BMC Evol Biol. 2011. PMID: 21473772 Free PMC article.
References
-
- Bird A. Perceptions of epigenetics. Nature. 2007;447:396–398. - PubMed
-
- Barrera LO, Ren B. The transcriptional regulatory code of eukaryotic cells - insights from genome-wide analysis of chromatin organization and transcription factor binding. Curr Opin Cell Biol. 2006;18:291–298. - PubMed
-
- Berger SL. The complex language of chromatin regulation during transcription. Nature. 2007;447:407–412. - PubMed
-
- Li B, Carey M, Workman JL. The role of chromatin during transcription. Cell. 2007;128:707–719. - PubMed
-
- Kouzarides T. Chromatin modifications and their function. Cell. 2007;128:693–705. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

