Meprins, membrane-bound and secreted astacin metalloproteinases
- PMID: 18783725
- PMCID: PMC2650038
- DOI: 10.1016/j.mam.2008.08.002
Meprins, membrane-bound and secreted astacin metalloproteinases
Abstract
The astacins are a subfamily of the metzincin superfamily of metalloproteinases. The first to be characterized was the crayfish enzyme astacin. To date more than 200 members of this family have been identified in species ranging from bacteria to humans. Astacins are involved in developmental morphogenesis, matrix assembly, tissue differentiation and digestion. Family members include the procollagen C-proteinase (BMP1, bone morphogenetic protein 1), tolloid and mammalian tolloid-like, HMP (Hydra vulgaris metalloproteinase), sea urchin BP10 (blastula protein) and SPAN (Strongylocentrotus purpuratus astacin), the 'hatching' subfamily comprising alveolin, ovastacin, LCE, HCE ('low' and 'high' choriolytic enzymes), nephrosin (from carp head kidney), UVS.2 from frog, and the meprins. In the human and mouse genomes, there are six astacin family genes (two meprins, three BMP1/tolloid-like, one ovastacin), but in Caenorhabditis elegans there are 40. Meprins are the only astacin proteinases that function on the membrane and extracellularly by virtue of the fact that they can be membrane-bound or secreted. They are unique in their domain structure and covalent subunit dimerization, oligomerization propensities, and expression patterns. They are normally highly regulated at the transcriptional and post-translational levels, localize to specific membranes or extracellular spaces, and can hydrolyse biologically active peptides, cytokines, extracellular matrix (ECM) proteins and cell-surface proteins. The in vivo substrates of meprins are unknown, but the abundant expression of these proteinases in the epithelial cells of the intestine, kidney and skin provide clues to their functions.
Figures
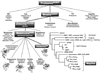


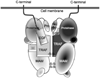


 5’ untranslated region;
5’ untranslated region;  signal peptide;
signal peptide;  propeptide; proteinase;
propeptide; proteinase;  MAM (meprin subunit domain/A5 protein/receptor protein tyrosine phosphatase l domain);
MAM (meprin subunit domain/A5 protein/receptor protein tyrosine phosphatase l domain);  MATH (meprin- and tumour-necrosis-factor-receptor-associated factors homology domain);
MATH (meprin- and tumour-necrosis-factor-receptor-associated factors homology domain);  intervening domain;
intervening domain;  I-domain (inserted domain);
I-domain (inserted domain);  EGF-like domain;
EGF-like domain;  transmembrane domain;
transmembrane domain;  cytosolic domain;
cytosolic domain;  3’ untranslated region (Hahn, Illisson et al. 2000).
3’ untranslated region (Hahn, Illisson et al. 2000).
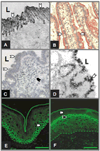
References
-
- Ambort D, Stalder D, Lottaz D, Hguenin M, Oneda B, Heller M, Sterchi EE. A novel 2D-based approach to the discovery of candidate substrates for the metalloendopeptidase meprin. FEBS J. 2008 - PubMed
-
- Bankus JM, Bond JS. Expression and distribution of meprin protease subunits in mouse intestine. Arch. Biochem. Biophys. 1996;331(1):87–94. - PubMed
-
- Barrett AJ, Rawlings ND, O'Brien EA. The MEROPS database as a protease information system. J. Struct. Biol. 2001;134(2–3):95–102. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

