Assessing the effect of varying sequence length on DNA barcoding of fungi
- PMID: 18784789
- PMCID: PMC1890918
- DOI: 10.1111/j.1471-8286.2007.01698.x
Assessing the effect of varying sequence length on DNA barcoding of fungi
Abstract
DNA barcoding shows enormous promise for the rapid identification of organisms at the species level. There has been much recent debate, however, about the need for longer barcode sequences, especially when these sequences are used to construct molecular phylogenies. Here, we have analysed a set of fungal mitochondrial sequences - of various lengths - and we have monitored the effect of reducing sequence length on the utility of the data for both species identification and phylogenetic reconstruction. Our results demonstrate that reducing sequence length has a profound effect on the accuracy of resulting phylogenetic trees, but surprisingly short sequences still yield accurate species identifications. We conclude that the standard short barcode sequences ( approximately 600 bp) are not suitable for inferring accurate phylogenetic relationships, but they are sufficient for species identification among the fungi.
Figures
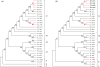
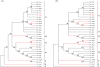



Similar articles
-
Characterizing the ribosomal tandem repeat and its utility as a DNA barcode in lichen-forming fungi.BMC Evol Biol. 2020 Jan 6;20(1):2. doi: 10.1186/s12862-019-1571-4. BMC Evol Biol. 2020. PMID: 31906844 Free PMC article.
-
Unambiguous identification of fungi: where do we stand and how accurate and precise is fungal DNA barcoding?IMA Fungus. 2020 Jul 10;11:14. doi: 10.1186/s43008-020-00033-z. eCollection 2020. IMA Fungus. 2020. PMID: 32714773 Free PMC article.
-
Highlights of DNA Barcoding in identification of salient microorganisms like fungi.J Mycol Med. 2016 Dec;26(4):291-297. doi: 10.1016/j.mycmed.2016.05.006. Epub 2016 Jul 9. J Mycol Med. 2016. PMID: 27402509 Review.
-
[Research progress on DNA barcoding analysis methods].Ying Yong Sheng Tai Xue Bao. 2018 Mar;29(3):1006-1014. doi: 10.13287/j.1001-9332.201803.032. Ying Yong Sheng Tai Xue Bao. 2018. PMID: 29722246 Chinese.
-
Fungal DNA barcoding.Genome. 2016 Nov;59(11):913-932. doi: 10.1139/gen-2016-0046. Epub 2016 Aug 30. Genome. 2016. PMID: 27829306 Review.
Cited by
-
Colletotrichum - current status and future directions.Stud Mycol. 2012 Sep 15;73(1):181-213. doi: 10.3114/sim0014. Stud Mycol. 2012. PMID: 23136460 Free PMC article.
-
Validation of the ITS2 region as a novel DNA barcode for identifying medicinal plant species.PLoS One. 2010 Jan 7;5(1):e8613. doi: 10.1371/journal.pone.0008613. PLoS One. 2010. PMID: 20062805 Free PMC article.
-
Multilocus Sequence Typing (MLST) and Random Polymorphic DNA (RAPD) Comparisons of Geographic Isolates of Neoparamoeba perurans, the Causative Agent of Amoebic Gill Disease.Pathogens. 2019 Nov 19;8(4):244. doi: 10.3390/pathogens8040244. Pathogens. 2019. PMID: 31752364 Free PMC article.
-
HACSim: an R package to estimate intraspecific sample sizes for genetic diversity assessment using haplotype accumulation curves.PeerJ Comput Sci. 2020 Jan 6;6:e243. doi: 10.7717/peerj-cs.243. eCollection 2020. PeerJ Comput Sci. 2020. PMID: 33816897 Free PMC article.
-
Towards barcode markers in Fungi: an intron map of Ascomycota mitochondria.BMC Bioinformatics. 2009 Jun 16;10 Suppl 6(Suppl 6):S15. doi: 10.1186/1471-2105-10-S6-S15. BMC Bioinformatics. 2009. PMID: 19534740 Free PMC article.
References
-
- Bullerwell CE, Leigh J, Seif E, Longcore JE, Lang BF. Evolution of the fungi and their mitochondrial genomes. In: Arora DK, Khachatourians GG, editors. Applied Mycology and Biotechnology. Vol. 3. Amsterdam: Elsevier Science; 2003b. pp. 133–159. Vol. 3 Fungal Genomics.
-
- Foster PG, Hickey DA. Compositional bias may affect both DNA-based and protein-based phylogenetic reconstructions. Journal of Molecular Evolution. 1999;48:284–290. - PubMed
LinkOut - more resources
Full Text Sources
