Sub-telomere directed gene expression during initiation of invasive aspergillosis
- PMID: 18787699
- PMCID: PMC2526178
- DOI: 10.1371/journal.ppat.1000154
Sub-telomere directed gene expression during initiation of invasive aspergillosis
Abstract
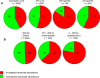
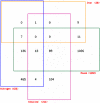
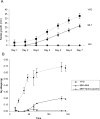
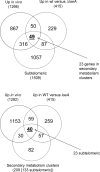
Aspergillus fumigatus is a common mould whose spores are a component of the normal airborne flora. Immune dysfunction permits developmental growth of inhaled spores in the human lung causing aspergillosis, a significant threat to human health in the form of allergic, and life-threatening invasive infections. The success of A. fumigatus as a pathogen is unique among close phylogenetic relatives and is poorly characterised at the molecular level. Recent genome sequencing of several Aspergillus species provides an exceptional opportunity to analyse fungal virulence attributes within a genomic and evolutionary context. To identify genes preferentially expressed during adaptation to the mammalian host niche, we generated multiple gene expression profiles from minute samplings of A. fumigatus germlings during initiation of murine infection. They reveal a highly co-ordinated A. fumigatus gene expression programme, governing metabolic and physiological adaptation, which allows the organism to prosper within the mammalian niche. As functions of phylogenetic conservation and genetic locus, 28% and 30%, respectively, of the A. fumigatus subtelomeric and lineage-specific gene repertoires are induced relative to laboratory culture, and physically clustered genes including loci directing pseurotin, gliotoxin and siderophore biosyntheses are a prominent feature. Locationally biased A. fumigatus gene expression is not prompted by in vitro iron limitation, acid, alkaline, anaerobic or oxidative stress. However, subtelomeric gene expression is favoured following ex vivo neutrophil exposure and in comparative analyses of richly and poorly nourished laboratory cultured germlings. We found remarkable concordance between the A. fumigatus host-adaptation transcriptome and those resulting from in vitro iron depletion, alkaline shift, nitrogen starvation and loss of the methyltransferase LaeA. This first transcriptional snapshot of a fungal genome during initiation of mammalian infection provides the global perspective required to direct much-needed diagnostic and therapeutic strategies and reveals genome organisation and subtelomeric diversity as potential driving forces in the evolution of pathogenicity in the genus Aspergillus.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




References
-
- Tekaia F, Latge JP. Aspergillus fumigatus: saprophyte or pathogen? Curr Opin Microbiol. 2005;8:385–392. - PubMed
-
- Zmeili OS, Soubani AO. Pulmonary aspergillosis: a clinical update. QJM. 2007;100:317–334. - PubMed
-
- Schaffner A, Douglas H, Braude A. Selective protection against conidia by mononuclear and against mycelia by polymorphonuclear phagocytes in resistance to Aspergillus. Observations on these two lines of defense in vivo and in vitro with human and mouse phagocytes. J Clin Invest. 1982;69:617–631. - PMC - PubMed
-
- Gersuk GM, Underhill DM, Zhu L, Marr KA. Dectin-1 and TLRs permit macrophages to distinguish between different Aspergillus fumigatus cellular states. J Immunol. 2006;176:3717–3724. - PubMed
Publication types
MeSH terms
Grants and funding
- BBS/B/10331/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- G0501164/MRC_/Medical Research Council/United Kingdom
- AI051144/AI/NIAID NIH HHS/United States
- R21 AI052236/AI/NIAID NIH HHS/United States
- G0501397/MRC_/Medical Research Council/United Kingdom
- R01 AI051144/AI/NIAID NIH HHS/United States
- N01 AI030041/AI/NIAID NIH HHS/United States
- P 18606/FWF_/Austrian Science Fund FWF/Austria
- WT_/Wellcome Trust/United Kingdom
- P30 CA016672/CA/NCI NIH HHS/United States
- AI052236/AI/NIAID NIH HHS/United States
- CA16672/CA/NCI NIH HHS/United States
- U01AI48830/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

