OSTM1 bone defect reveals an intercellular hematopoietic crosstalk
- PMID: 18790735
- PMCID: PMC2662145
- DOI: 10.1074/jbc.M805242200
OSTM1 bone defect reveals an intercellular hematopoietic crosstalk
Abstract
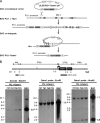
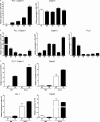
The most severe form of bone autosomal recessive osteopetrosis both in humans and in the gray-lethal (gl/gl) mouse is caused by mutations in the Ostm1 gene. Although osteopetrosis is usually associated with a defect in the hematopoietic-derived osteoclast cells, this study determined that Ostm1 is expressed in many hematopoietic cells of the myeloid and lymphoid B- and T-lineages. Hematopoiesis in gl/gl mice is characterized by a marked expansion of the osteoclast lineage but also by deregulation of the lymphoid lineages with a decrease in B-lymphoid cell populations and altered distribution in T-lymphoid double and single CD4 CD8-positive cells. In committed gl/gl osteoclasts, specific Ostm1 transgene targeting showed a requirement of additional factors and/or cells for normal osteoclast function, and importantly, defined the gl osteopetrotic defect as non-cell autonomous. By contrast, gl/gl osteoclast, B- and T-lymphoid lineage phenotypes were rescued when Ostm1 is expressed under PU.1 regulation from a bacterial artificial chromosome transgene, which established an essential role for Ostm1 in hematopoietic cells in addition to osteoclasts. Together these experiments are the first to demonstrate the existence of hematopoietic crosstalk for the production of functional and active osteoclasts.
Figures



References
-
- Teitelbaum, S. L., and Ross, P. (2003) Nat. Rev. Genet. 4 638–649 - PubMed
-
- Boyle, W. J., Simonet, W. S., and Lacey, D. L. (2003) Nature 423 337–342 - PubMed
-
- Zaidi, M. (2007) Nat. Med. 13 791–801 - PubMed
-
- Janssens, K., and Van Hul, W. (2002) Hum. Mol. Genet. 11 2385–2393 - PubMed
-
- Lazner, F., Gowen, M., Pavasovic, D., and Kola, I. (1999) Hum. Mol. Genet. 8 1839–1846 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

