A genome-scale metabolic reconstruction of Pseudomonas putida KT2440: iJN746 as a cell factory
- PMID: 18793442
- PMCID: PMC2569920
- DOI: 10.1186/1752-0509-2-79
A genome-scale metabolic reconstruction of Pseudomonas putida KT2440: iJN746 as a cell factory
Abstract
Background: Pseudomonas putida is the best studied pollutant degradative bacteria and is harnessed by industrial biotechnology to synthesize fine chemicals. Since the publication of P. putida KT2440's genome, some in silico analyses of its metabolic and biotechnology capacities have been published. However, global understanding of the capabilities of P. putida KT2440 requires the construction of a metabolic model that enables the integration of classical experimental data along with genomic and high-throughput data. The constraint-based reconstruction and analysis (COBRA) approach has been successfully used to build and analyze in silico genome-scale metabolic reconstructions.
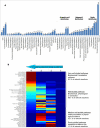
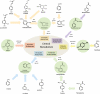
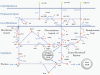
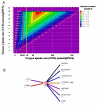
Results: We present a genome-scale reconstruction of P. putida KT2440's metabolism, iJN746, which was constructed based on genomic, biochemical, and physiological information. This manually-curated reconstruction accounts for 746 genes, 950 reactions, and 911 metabolites. iJN746 captures biotechnologically relevant pathways, including polyhydroxyalkanoate synthesis and catabolic pathways of aromatic compounds (e.g., toluene, benzoate, phenylacetate, nicotinate), not described in other metabolic reconstructions or biochemical databases. The predictive potential of iJN746 was validated using experimental data including growth performance and gene deletion studies. Furthermore, in silico growth on toluene was found to be oxygen-limited, suggesting the existence of oxygen-efficient pathways not yet annotated in P. putida's genome. Moreover, we evaluated the production efficiency of polyhydroxyalkanoates from various carbon sources and found fatty acids as the most prominent candidates, as expected.
Conclusion: Here we presented the first genome-scale reconstruction of P. putida, a biotechnologically interesting all-surrounder. Taken together, this work illustrates the utility of iJN746 as i) a knowledge-base, ii) a discovery tool, and iii) an engineering platform to explore P. putida's potential in bioremediation and bioplastic production.
Figures
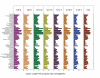
References
-
- Clarke P, Richmond MH. Genetics and Biochemistry of Pseudomonas. New York, USA: John Wiley & Sons; 1975.
-
- Franklin FC, Bagdasarian M, Bagdasarian MM, Timmis K. Molecular and functional analysis of the TOL plasmid pWWO from Pseudomonas putida and cloning of genes for the entire regulated aromatic ring meta cleavage pathway. Proc Natl Acad Sci USA. 1981;78:7458–7462. doi: 10.1073/pnas.78.12.7458. - DOI - PMC - PubMed
-
- Mermod N, Harayama S, Timmis K. New route to bacterial production of indigo. Bio/Technology. 1986;4:321–324. doi: 10.1038/nbt0486-321. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

