Origin and pathogenesis of nodular lymphocyte-predominant Hodgkin lymphoma as revealed by global gene expression analysis
- PMID: 18794340
- PMCID: PMC2556780
- DOI: 10.1084/jem.20080809
Origin and pathogenesis of nodular lymphocyte-predominant Hodgkin lymphoma as revealed by global gene expression analysis
Abstract
The pathogenesis of nodular lymphocyte-predominant Hodgkin lymphoma (NLPHL) and its relationship to other lymphomas are largely unknown. This is partly because of the technical challenge of analyzing its rare neoplastic lymphocytic and histiocytic (L&H) cells, which are dispersed in an abundant nonneoplastic cellular microenvironment. We performed a genome-wide expression study of microdissected L&H lymphoma cells in comparison to normal and other malignant B cells that indicated a relationship of L&H cells to and/or that they originate from germinal center B cells at the transition to memory B cells. L&H cells show a surprisingly high similarity to the tumor cells of T cell-rich B cell lymphoma and classical Hodgkin lymphoma, a partial loss of their B cell phenotype, and deregulation of many apoptosis regulators and putative oncogenes. Importantly, L&H cells are characterized by constitutive nuclear factor kappaB activity and aberrant extracellular signal-regulated kinase signaling. Thus, these findings shed new light on the nature of L&H cells, reveal several novel pathogenetic mechanisms in NLPHL, and may help in differential diagnosis and lead to novel therapeutic strategies.
Figures

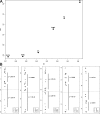



References
-
- Küppers, R. 2002. Molecular biology of Hodgkin's lymphoma. Adv. Cancer Res. 84:277–312. - PubMed
-
- Schwering, I., A. Bräuninger, U. Klein, B. Jungnickel, M. Tinguely, V. Diehl, M.L. Hansmann, R. Dalla-Favera, K. Rajewsky, and R. Küppers. 2003. Loss of the B-lineage-specific gene expression program in Hodgkin and Reed-Sternberg cells of Hodgkin lymphoma. Blood. 101:1505–1512. - PubMed
-
- Falini, B., B. Bigerna, L. Pasqualucci, M. Fizzotti, M.F. Martelli, S. Pileri, A. Pinto, A. Carbone, S. Venturi, R. Pacini, et al. 1996. Distinctive expression pattern of the BCL-6 protein in nodular lymphocyte predominance Hodgkin's disease. Blood. 87:465–471. - PubMed
-
- Greiner, A., S. Tobollik, M. Buettner, B. Jungnickel, K. Herrmann, E. Kremmer, and G. Niedobitek. 2005. Differential expression of activation-induced cytidine deaminase (AID) in nodular lymphocyte-predominant and classical Hodgkin lymphoma. J. Pathol. 205:541–547. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

