Mutational analysis of vaccinia virus mRNA cap (guanine-N7) methyltransferase reveals essential contributions of the N-terminal peptide that closes over the active site
- PMID: 18799596
- PMCID: PMC2578867
- DOI: 10.1261/rna.1201308
Mutational analysis of vaccinia virus mRNA cap (guanine-N7) methyltransferase reveals essential contributions of the N-terminal peptide that closes over the active site
Abstract
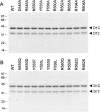
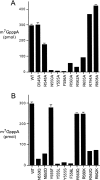
RNA guanine-N7 methyltransferase catalyzes the third step of eukaryal mRNA capping, the transfer of a methyl group from AdoMet to GpppRNA to form m(7)GpppRNA. Mutational and crystallographic analyses of cellular and poxvirus cap methyltransferases have yielded a coherent picture of a conserved active site and determinants of substrate specificity. Models of the Michaelis complex suggest a direct in-line mechanism of methyl transfer. Because no protein contacts to the guanine-N7 nucleophile, the AdoMet methyl carbon (Cepsilon) or the AdoHcy sulfur (Sdelta) leaving group were observed in ligand-bound structures of cellular cap methyltransferase, it was initially thought that the enzyme facilitates catalysis by optimizing proximity and geometry of the donor and acceptor. However, the structure of AdoHcy-bound vaccinia virus cap methyltransferase revealed the presence of an N-terminal "lid peptide" that closes over the active site and makes multiple contacts with the substrates, including the AdoMet sulfonium. This segment is disordered in the vaccinia apoenzyme and is not visible in the available structures of cellular cap methyltransferase. Here, we conducted a mutational analysis of the vaccinia virus lid peptide ((545)DKFRLNPEVSYFTNKRTRG(563)) entailing in vivo and in vitro readouts of the effects of alanine and conservative substitutions. We thereby identified essential functional groups that interact with the AdoMet sulfonium (Tyr555, Phe556), the AdoMet adenine (Asn550), and the cap triphosphate bridge (Arg560, Arg562). The results suggest that van der Waals contacts of Tyr555 and Phe556 to the AdoMet Sdelta and C epsilon atoms, and the electron-rich environment around the sulfonium, serve to stabilize the transition state of the transmethylation reaction.
Figures

References
-
- Bujnicki J.M., Feder M., Radlinska M., Rychlewski L. mRNA:guanine-N7 methyltransferases: Identification of novel members of the family, evolutionary analysis, homology modeling, and analysis of sequence-structure-function relationships. BMC Bioinformatics. 2001;2:2. doi: 10.1186/1471-2105-2-2. - DOI - PMC - PubMed
-
- Cong P., Shuman S. Methyltransferase and subunit association domains of vaccinia virus mRNA capping enzyme. J. Biol. Chem. 1992;267:16424–16429. - PubMed
-
- Fabrega C., Hausmann S., Shen V., Shuman S., Lima C.D. Structure and mechanism of cap (guanine-N7) methyltransferase. Mol. Cell. 2004;13:77–89. - PubMed
-
- Hausmann S., Zheng S., Fabrega C., Schneller S.W., Lima C.D., Shuman S. Encephalitozoon cuniculi mRNA cap (guanine-N7) methyltransferase: Methyl acceptor specificity, inhibition by AdoMet analogs, and structure-guided mutational analysis. J. Biol. Chem. 2005;280:20404–20412. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
