Exon creation and establishment in human genes
- PMID: 18811936
- PMCID: PMC2592719
- DOI: 10.1186/gb-2008-9-9-r141
Exon creation and establishment in human genes
Abstract
Background: A large proportion of species-specific exons are alternatively spliced. In primates, Alu elements play a crucial role in the process of exon creation but many new exons have appeared through other mechanisms. Despite many recent studies, it is still unclear which are the splicing regulatory requirements for de novo exonization and how splicing regulation changes throughout an exon's lifespan.
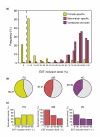
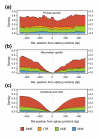
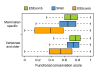
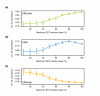
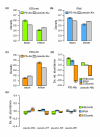
Results: Using comparative genomics, we have defined sets of exons with different evolutionary ages. Younger exons have weaker splice-sites and lower absolute values for the relative abundance of putative splicing regulators between exonic and adjacent intronic regions, indicating a less consolidated splicing regulation. This relative abundance is shown to increase with exon age, leading to higher exon inclusion. We show that this local difference in the density of regulators might be of biological significance, as it outperforms other measures in real exon versus pseudo-exon classification. We apply this new measure to the specific case of the exonization of anti-sense Alu elements and show that they are characterized by a general lack of exonic splicing silencers.
Conclusions: Our results suggest that specific sequence environments are required for exonization and that these can change with time. We propose a model of exon creation and establishment in human genes, in which splicing decisions depend on the relative local abundance of regulatory motifs. Using this model, we provide further explanation as to why Alu elements serve as a major substrate for exon creation in primates. Finally, we discuss the benefits of integrating such information in gene prediction.
Figures

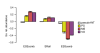
Similar articles
-
RNA-editing-mediated exon evolution.Genome Biol. 2007;8(2):R29. doi: 10.1186/gb-2007-8-2-r29. Genome Biol. 2007. PMID: 17326827 Free PMC article.
-
Alternative splicing of Alu exons--two arms are better than one.Nucleic Acids Res. 2008 Apr;36(6):2012-23. doi: 10.1093/nar/gkn024. Epub 2008 Feb 14. Nucleic Acids Res. 2008. PMID: 18276646 Free PMC article.
-
Exonization of transposed elements: A challenge and opportunity for evolution.Biochimie. 2011 Nov;93(11):1928-34. doi: 10.1016/j.biochi.2011.07.014. Epub 2011 Jul 26. Biochimie. 2011. PMID: 21787833 Review.
-
Diverse splicing patterns of exonized Alu elements in human tissues.PLoS Genet. 2008 Oct 17;4(10):e1000225. doi: 10.1371/journal.pgen.1000225. PLoS Genet. 2008. PMID: 18841251 Free PMC article.
-
The birth of new exons: mechanisms and evolutionary consequences.RNA. 2007 Oct;13(10):1603-8. doi: 10.1261/rna.682507. Epub 2007 Aug 20. RNA. 2007. PMID: 17709368 Free PMC article. Review.
Cited by
-
Transcriptome-wide expansion of non-coding regulatory switches: evidence from co-occurrence of Alu exonization, antisense and editing.Nucleic Acids Res. 2013 Feb 1;41(4):2121-37. doi: 10.1093/nar/gks1457. Epub 2013 Jan 8. Nucleic Acids Res. 2013. PMID: 23303787 Free PMC article.
-
Changes in exon-intron structure during vertebrate evolution affect the splicing pattern of exons.Genome Res. 2012 Jan;22(1):35-50. doi: 10.1101/gr.119834.110. Epub 2011 Oct 5. Genome Res. 2012. PMID: 21974994 Free PMC article.
-
Sequence composition similarities with the 7SL RNA are highly predictive of functional genomic features.Nucleic Acids Res. 2010 Aug;38(15):4907-16. doi: 10.1093/nar/gkq234. Epub 2010 Apr 14. Nucleic Acids Res. 2010. PMID: 20392819 Free PMC article.
-
Novel kinin B₁ receptor splice variant and 5'UTR regulatory elements are responsible for cell specific B₁ receptor expression.PLoS One. 2014 Jan 27;9(1):e87175. doi: 10.1371/journal.pone.0087175. eCollection 2014. PLoS One. 2014. PMID: 24475248 Free PMC article.
-
Deep intronic mutations and human disease.Hum Genet. 2017 Sep;136(9):1093-1111. doi: 10.1007/s00439-017-1809-4. Epub 2017 May 12. Hum Genet. 2017. PMID: 28497172 Review.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

