A familial inverted duplication/deletion of 2p25.1-25.3 provides new clues on the genesis of inverted duplications
- PMID: 18813332
- PMCID: PMC2986066
- DOI: 10.1038/ejhg.2008.160
A familial inverted duplication/deletion of 2p25.1-25.3 provides new clues on the genesis of inverted duplications
Abstract
We studied a family in which the same 10 Mb inverted duplication of 2p25.3-p25.1 segregates in two children and their father, all showing a trisomy phenotype. As FISH analysis demonstrated that the duplication was inverted, we suspected that a contiguous terminal deletion was also present, according to the classical inv dup del type of rearrangements. Although FISH with 2p and 2q subtelomeric probes gave normal results, 100 kb resolution array-C/GH (aCGH) showed that, beside the duplication, a 273 kb deletion was also present. The presence of a single-copy region between the deleted and duplicated regions was further suspected through high-resolution aCGH analysis (approximately 20 kb), although only one informative spot having a normal log ratio was detected. The precise structure of the rearrangement was re-defined by real-time PCR and breakpoint cloning, demonstrating the presence of a 2680 bp single-copy sequence between deleted and duplicated regions and the involvement of a simple repeat with the potential for forming a non-B DNA structure. The rearrangement was not mediated by segmental duplications or short inverted repeats, and the double-strand break might have been repaired by non-homologous end joining or microhomology-mediated intrastrand repair. These data highlight the fact that concomitant deletions associated with inverted duplications are very likely to be more frequent than classical cytogenetic methods alone have been able to demonstrate. The phenotypic effects of the trisomy and of the terminal 2p deletion are discussed.
Figures
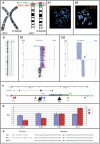

References
-
- Phelan MC, Crawford EC, Bealer DM. Mental retardation in South Carolina III. Chromosome aberrations. Proc Greenwood Genet Center. 1996;15:45–60.
-
- Shaffer LG, Lupski JR. Molecular mechanisms for constitutional chromosomal rearrangements in humans. Annu Rev Genet. 2000;34:297–329. - PubMed
-
- Van Dyke DL, Miller MJ, Weiss L. The origin of inverted tandem duplications, and phenotypic effects of tandem duplication of the X chromosome long arm. Am J Med Genet. 1983;15:441–450. - PubMed
-
- Shaw CJ, Lupski JR. Implications of human genome architecture for rearrangement-based disorders: the genomic basis of disease. Hum Mol Genet. 2004;13:R57–R64. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

