The Apicomplexan whole-genome phylogeny: an analysis of incongruence among gene trees
- PMID: 18820254
- PMCID: PMC2582981
- DOI: 10.1093/molbev/msn213
The Apicomplexan whole-genome phylogeny: an analysis of incongruence among gene trees
Abstract
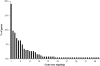
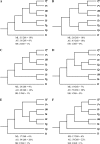
The protistan phylum Apicomplexa contains many important pathogens and is the subject of intense genome sequencing efforts. Based upon the genome sequences from seven apicomplexan species and a ciliate outgroup, we identified 268 single-copy genes suitable for phylogenetic inference. Both concatenation and consensus approaches inferred the same species tree topology. This topology is consistent with most prior conceptions of apicomplexan evolution based upon ultrastructural and developmental characters, that is, the piroplasm genera Theileria and Babesia form the sister group to the Plasmodium species, the coccidian genera Eimeria and Toxoplasma are monophyletic and are the sister group to the Plasmodium species and piroplasm genera, and Cryptosporidium forms the sister group to the above mentioned with the ciliate Tetrahymena as the outgroup. The level of incongruence among gene trees appears to be high at first glance; only 19% of the genes support the species tree, and a total of 48 different gene-tree topologies are observed. Detailed investigations suggest that the low signal-to-noise ratio in many genes may be the main source of incongruence. The probability of being consistent with the species tree increases as a function of the minimum bootstrap support observed at tree nodes for a given gene tree. Moreover, gene sequences that generate high bootstrap support are robust to the changes in alignment parameters or phylogenetic method used. However, caution should be taken in that some genes can infer a "wrong" tree with strong support because of paralogy, model violations, or other causes. The importance of examining multiple, unlinked genes that possess a strong phylogenetic signal cannot be overstated.
Figures

References
-
- Abrahamsen MS, Templeton TJ, Enomoto S, et al. (20 co-authors) Complete genome sequence of the apicomplexan, Cryptosporidium parvum. Science. 2004;304:441–445. - PubMed
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. - PubMed
-
- Bergsten J. A review of long-branch attraction. Cladistics. 2005;21:163–193. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

