High-throughput Agrobacterium-mediated barley transformation
- PMID: 18822125
- PMCID: PMC2562381
- DOI: 10.1186/1746-4811-4-22
High-throughput Agrobacterium-mediated barley transformation
Abstract
Background: Plant transformation is an invaluable tool for basic plant research, as well as a useful technique for the direct improvement of commercial crops. Barley (Hordeum vulgare) is the fourth most abundant cereal crop in the world. It also provides a useful model for the study of wheat, which has a larger and more complex genome. Most existing barley transformation methodologies are either complex or have low (<10%) transformation efficiencies.
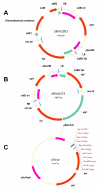
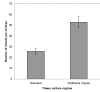
Results: A robust, simple and reproducible barley transformation protocol has been developed that yields average transformation efficiencies of 25%. This protocol is based on the infection of immature barley embryos with Agrobacterium strain AGL1, carrying vectors from the pBract series that contain the hpt gene (conferring hygromycin resistance) as a selectable marker. Results of large scale experiments utilising the luc (firefly luciferase) gene as a reporter are described. The method presented here has been used to produce hundreds of independent, transgenic plant lines and we show that a large proportion of these lines contain single copies of the luc gene.
Conclusion: This protocol demonstrates significant improvements in both efficiency and ease of use over existing barley transformation methods. This opens up opportunities for the development of functional genomics resources in barley.
Figures



References
-
- Tingay S, McElroy D, Kalla R, Fieg S, Wang M, Thornton S, Brettell R. Agrobacterium tumefaciens-mediated barley transformation. Plant J. 1997;11:1369–1376. doi: 10.1046/j.1365-313X.1997.11061369.x. - DOI
-
- Zhang S, Cho MJ, Koprek T, Yun R, Bregitzer P, Lemaux PG. Genetic transformation of commercial cultivars of oat (Avena sativa L.) and barley (Hordeum vulgare L.) using shoot meristematic cultures derived from germinated seedlings. Plant Cell Rep. 1999;18:959–966. doi: 10.1007/s002990050691. - DOI
LinkOut - more resources
Full Text Sources
Other Literature Sources

