Three-dimensional structures of Pseudomonas aeruginosa PvcA and PvcB, two proteins involved in the synthesis of 2-isocyano-6,7-dihydroxycoumarin
- PMID: 18824174
- PMCID: PMC2701257
- DOI: 10.1016/j.jmb.2008.09.027
Three-dimensional structures of Pseudomonas aeruginosa PvcA and PvcB, two proteins involved in the synthesis of 2-isocyano-6,7-dihydroxycoumarin
Abstract
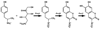
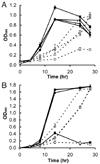
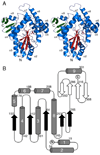
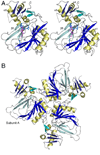
The pvcABCD operon of Pseudomonas aeruginosa encodes four proteins (PA2254, PA2255, PA2256, and PA2257) that form a cluster that is responsible for the synthesis of a cyclized isocyano derivative of tyrosine. These proteins, which were identified originally as being responsible for a step in the maturation of the chromophore of the peptide siderophore pyoverdine, have been identified recently as belonging to a family of proteins that produce small organic isonitriles. We report that strains harboring a disruption in the pvcA or pvcB genes are able to grow in iron-depleted conditions and to produce pyoverdine. Additionally, we have determined the three-dimensional crystal structures of PvcA and PvcB. The structure of PvcA demonstrates a novel enzyme architecture that is built upon a Rossmann fold. We have analyzed the sequence conservation of enzymes within this family and identified six conserved motifs. These regions of the protein cluster around a putative active site cavity. The structure of the PvcB protein confirms it is a member of the Fe2+/alpha-ketoglutarate-dependent oxygenase family of enzymes. The active site of PvcB is compared to the structures of other family members and suggests that a conformational change to order several loops will accompany the binding of ligands.
Figures


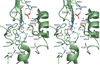
Similar articles
-
The 1.8 A crystal structure of PA2412, an MbtH-like protein from the pyoverdine cluster of Pseudomonas aeruginosa.J Biol Chem. 2007 Jul 13;282(28):20425-34. doi: 10.1074/jbc.M611833200. Epub 2007 May 14. J Biol Chem. 2007. PMID: 17502378
-
Examining Reaction Specificity in PvcB, a Source of Diversity in Isonitrile-Containing Natural Products.Biochemistry. 2015 Apr 28;54(16):2659-69. doi: 10.1021/acs.biochem.5b00255. Epub 2015 Apr 20. Biochemistry. 2015. PMID: 25866990
-
The Pseudomonas aeruginosa extracellular secondary metabolite, Paerucumarin, chelates iron and is not localized to extracellular membrane vesicles.J Microbiol. 2016 Aug;54(8):573-81. doi: 10.1007/s12275-016-5645-3. Epub 2016 Aug 2. J Microbiol. 2016. PMID: 27480638
-
The pvc gene cluster of Pseudomonas aeruginosa: role in synthesis of the pyoverdine chromophore and regulation by PtxR and PvdS.J Bacteriol. 1999 Jul;181(13):4118-24. doi: 10.1128/JB.181.13.4118-4124.1999. J Bacteriol. 1999. PMID: 10383985 Free PMC article.
-
Mutation in pvcABCD operon of Pseudomonas aeruginosa modulates MexEF-OprN efflux system and hence resistance to chloramphenicol and ciprofloxacin.Microb Pathog. 2020 Dec;149:104491. doi: 10.1016/j.micpath.2020.104491. Epub 2020 Sep 14. Microb Pathog. 2020. PMID: 32941967
Cited by
-
Distribution and prediction of catalytic domains in 2-oxoglutarate dependent dioxygenases.BMC Res Notes. 2012 Aug 4;5:410. doi: 10.1186/1756-0500-5-410. BMC Res Notes. 2012. PMID: 22862831 Free PMC article.
-
Pseudomonas aeruginosa L10: A Hydrocarbon-Degrading, Biosurfactant-Producing, and Plant-Growth-Promoting Endophytic Bacterium Isolated From a Reed (Phragmites australis).Front Microbiol. 2018 May 25;9:1087. doi: 10.3389/fmicb.2018.01087. eCollection 2018. Front Microbiol. 2018. PMID: 29887849 Free PMC article.
-
An iron detection system determines bacterial swarming initiation and biofilm formation.Sci Rep. 2016 Nov 15;6:36747. doi: 10.1038/srep36747. Sci Rep. 2016. PMID: 27845335 Free PMC article.
-
Actinobacterial chalkophores: the biosynthesis of hazimycins.J Antibiot (Tokyo). 2024 Apr;77(4):228-237. doi: 10.1038/s41429-024-00706-6. Epub 2024 Feb 20. J Antibiot (Tokyo). 2024. PMID: 38378905
-
Studies on the Properties of the Sporulation Specific Protein Dit1 and its Product Formyl Tyrosine.J Fungi (Basel). 2020 Jun 3;6(2):77. doi: 10.3390/jof6020077. J Fungi (Basel). 2020. PMID: 32503197 Free PMC article.
References
-
- Visca P, Imperi F, Lamont IL. Pyoverdine siderophores: from biogenesis to biosignificance. Trends Microbiol. 2007;15:22–30. - PubMed
-
- Sattely ES, Fischbach MA, Walsh CT. Total biosynthesis: in vitro reconstitution of polyketide and nonribosomal peptide pathways. Nat Prod Rep. 2008;25:757–793. - PubMed
-
- Weissman KJ, Muller R. Protein-protein interactions in multienzyme megasynthetases. Chembiochem. 2008;9:826–848. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

