A conserved domain of previously unknown function in Gap1 mediates protein-protein interaction and is required for biogenesis of a serine-rich streptococcal adhesin
- PMID: 18826412
- PMCID: PMC2938783
- DOI: 10.1111/j.1365-2958.2008.06456.x
A conserved domain of previously unknown function in Gap1 mediates protein-protein interaction and is required for biogenesis of a serine-rich streptococcal adhesin
Abstract
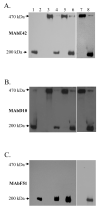
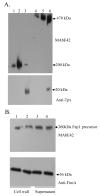
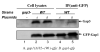
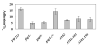
Fap1-like serine-rich proteins are a new family of bacterial adhesins found in a variety of streptococci and staphylococci that have been implicated in bacterial pathogenesis. A gene cluster encoding glycosyltransferases and accessory Sec components is required for Fap1 glycosylation and biogenesis in Streptococcus parasanguinis. Here we report that the glycosylation-associated protein, Gap1, contributes to glycosylation and biogenesis of Fap1 by interacting with another glycosylation-associated protein, Gap3. Gap1 shares structural homology with glycosyltransferases. The gap1 mutant, like the gap3 mutant, produced an aberrantly glycosylated Fap1 precursor and failed to produce mature Fap1, suggesting that Gap1 and Gap3 might function in concert in the Fap1 glycosylation and biogenesis. Indeed, Gap1 interacted with Gap3 in vitro and in vivo. A Gap1 N-terminal motif, within a highly conserved domain of unknown function (DUF1975) identified in many bacterial glycosyltransferases, was required for the Gap1-Gap3 interaction. Deletion of one, four and nine amino acids within the conserved motif gradually inhibited the Gap1-Gap3 interaction and diminished production of mature Fap1 and concurrently increased production of the Fap1 precursor. Consequently, bacterial adhesion to an in vitro tooth model was also reduced. These data demonstrate that the Gap1-Gap3 interaction is required for Fap1 biogenesis and Fap1-dependent bacterial adhesion.
Figures




Similar articles
-
Gap1 functions as a molecular chaperone to stabilize its interactive partner Gap3 during biogenesis of serine-rich repeat bacterial adhesin.Mol Microbiol. 2012 Feb;83(4):866-78. doi: 10.1111/j.1365-2958.2012.07970.x. Epub 2012 Jan 18. Mol Microbiol. 2012. PMID: 22251284 Free PMC article.
-
A conserved C-terminal 13-amino-acid motif of Gap1 is required for Gap1 function and necessary for the biogenesis of a serine-rich glycoprotein of Streptococcus parasanguinis.Infect Immun. 2008 Dec;76(12):5624-31. doi: 10.1128/IAI.00534-08. Epub 2008 Oct 13. Infect Immun. 2008. PMID: 18852249 Free PMC article.
-
Canonical SecA associates with an accessory secretory protein complex involved in biogenesis of a streptococcal serine-rich repeat glycoprotein.J Bacteriol. 2011 Dec;193(23):6560-6. doi: 10.1128/JB.05668-11. Epub 2011 Sep 30. J Bacteriol. 2011. PMID: 21965576 Free PMC article.
-
Glycosylation and biogenesis of a family of serine-rich bacterial adhesins.Microbiology (Reading). 2009 Feb;155(Pt 2):317-327. doi: 10.1099/mic.0.025221-0. Microbiology (Reading). 2009. PMID: 19202081 Review.
-
Glycosyltransferase-mediated Sweet Modification in Oral Streptococci.J Dent Res. 2015 May;94(5):659-65. doi: 10.1177/0022034515574865. Epub 2015 Mar 9. J Dent Res. 2015. PMID: 25755271 Free PMC article. Review.
Cited by
-
Both GtfA and GtfB are required for SraP glycosylation in Staphylococcus aureus.Curr Microbiol. 2014 Aug;69(2):121-6. doi: 10.1007/s00284-014-0563-2. Curr Microbiol. 2014. PMID: 24658735
-
Unraveling the sequence of cytosolic reactions in the export of GspB adhesin from Streptococcus gordonii.J Biol Chem. 2018 Apr 6;293(14):5360-5373. doi: 10.1074/jbc.RA117.000963. Epub 2018 Feb 9. J Biol Chem. 2018. PMID: 29462788 Free PMC article.
-
Gap1 functions as a molecular chaperone to stabilize its interactive partner Gap3 during biogenesis of serine-rich repeat bacterial adhesin.Mol Microbiol. 2012 Feb;83(4):866-78. doi: 10.1111/j.1365-2958.2012.07970.x. Epub 2012 Jan 18. Mol Microbiol. 2012. PMID: 22251284 Free PMC article.
-
Structure of a novel O-linked N-acetyl-D-glucosamine (O-GlcNAc) transferase, GtfA, reveals insights into the glycosylation of pneumococcal serine-rich repeat adhesins.J Biol Chem. 2014 Jul 25;289(30):20898-907. doi: 10.1074/jbc.M114.581934. J Biol Chem. 2014. PMID: 24936067 Free PMC article.
-
A novel glucosyltransferase is required for glycosylation of a serine-rich adhesin and biofilm formation by Streptococcus parasanguinis.J Biol Chem. 2010 Apr 16;285(16):12140-8. doi: 10.1074/jbc.M109.066928. Epub 2010 Feb 17. J Biol Chem. 2010. PMID: 20164186 Free PMC article.
References
-
- Bennett-Lovsey RM, Herbert AD, Sternberg MJ, Kelley LA. Exploring the extremes of sequence/structure space with ensemble fold recognition in the program Phyre. Proteins. 2008;70:611–625. - PubMed
-
- Bensing BA, Sullam PM. An accessory sec locus of Streptococcus gordonii is required for export of the surface protein GspB and for normal levels of binding to human platelets. Mol Microbiol. 2002;44:1081–1094. - PubMed
-
- Carlsson J, Grahnen H, Jonsson G, Wikner S. Establishment of Streptococcus sanguis in the mouths of infants. Arch Oral Biol. 1970;15:1143–1148. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

