Transcriptome content and dynamics at single-nucleotide resolution
- PMID: 18828881
- PMCID: PMC2592708
- DOI: 10.1186/gb-2008-9-9-234
Transcriptome content and dynamics at single-nucleotide resolution
Abstract
Massively parallel short-tag sequencing of cDNA libraries--RNAseq--is being used to study the dynamics and complexity of eukaryotic transcriptomes, giving new biological insights into the 'active genome'.
Figures
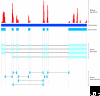
References
-
- Sultan M, Schulz MH, Richard H, Magen A, Klingenhoff A, Scherf M, Seifert M, Borodina T, Soldatov A, Parkhomchuk D, Schmidt D, O'Keeffe S, Haas S, Vingron M, Lehrach H, Yaspo M-L. A global view of gene activity and alternative splicing by deep sequencing of the human transcriptome. Science. 2008;321:956–960. doi: 10.1126/science.1160342. - DOI - PubMed
-
- Cloonan N, Forrest AR, Kolle G, Gardiner BBA, Faulkner GJ, Brown MK, Taylor DF, Steptoe AL, Wani S, Bethel G, Robertson AJ, Perkins AC, Bruce SJ, Lee CC, Ranade SS, Peckham HE, Manning JM, McKernan KJ, Grimmond SM. Stem cell transcriptome profiling via massive-scale mRNA sequencing. Nat Methods. 2008;5:613–619. doi: 10.1038/nmeth.1223. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

