Principles of transcriptional regulation and evolution of the metabolic system in E. coli
- PMID: 18836036
- PMCID: PMC2612968
- DOI: 10.1101/gr.079715.108
Principles of transcriptional regulation and evolution of the metabolic system in E. coli
Abstract
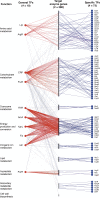
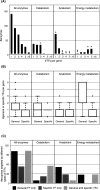
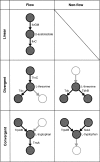
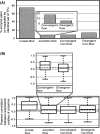
Organisms must adapt to make optimal use of the metabolic system in response to environmental changes. In the long-term, this involves evolution of the genomic repertoire of enzymes; in the short-term, transcriptional control ensures that appropriate enzymes are expressed in response to transitory extracellular conditions. Unicellular organisms are particularly susceptible to environmental changes; however, genome-scale impact of these modulatory effects has not been explored so far in bacteria. Here, we integrate genome-scale data to investigate the evolutionary trends and transcriptional control of metabolism in Escherichia coli K12. Globally, the regulatory system is organized in a clear hierarchy of general and specific transcription factors (TFs) that control differing ranges of metabolic functions. Further, catabolic, anabolic, and central metabolic pathways are targeted by distinct combinations of these TFs. Locally, enzymes catalyzing sequential reactions in a metabolic pathway are co-regulated by the same TFs. Regulation is more complex at junctions: General TFs control the overall activity of all connecting reactions, whereas specific TFs control individual enzymes. Divergent junctions play a special role in delineating metabolic pathways and decouple the regulation of incoming and outgoing reactions. We find little evidence for differential usage of isozymes, which are generally co-expressed in similar conditions, and thus are likely to reinforce the metabolic system through redundancy. Finally, we show that enzymes controlled by the same TFs have a strong tendency to co-evolve, suggesting a significant constraint to maintain similar regulatory regimes during evolution. Catabolic, anabolic, and central energy pathways evolve differently, emphasizing the role of the environment in shaping the metabolic system. Many of the observations also occur in yeast, and our findings may apply across large evolutionary distances.
Figures


References
-
- Almaas E., Kovacs B., Vicsek T., Oltvai Z.N., Barabasi A.L. Global organization of metabolic fluxes in the bacterium Escherichia coli. Nature. 2004;427:839–843. - PubMed
-
- Alon U. Network motifs: Theory and experimental approaches. Nat. Rev. Genet. 2007;8:450–461. - PubMed
-
- Al-Shahrour F., Diaz-Uriarte R., Dopazo J. FatiGO: A web tool for finding significant associations of gene ontology terms with groups of genes. Bioinformatics. 2004;20:578–580. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
