Probing the lower size limit for protein-like fold stability: ten-residue microproteins with specific, rigid structures in water
- PMID: 18842046
- PMCID: PMC2677142
- DOI: 10.1021/ja804656h
Probing the lower size limit for protein-like fold stability: ten-residue microproteins with specific, rigid structures in water
Abstract
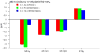
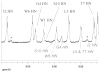
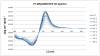
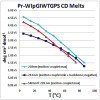
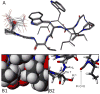
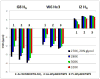
Mutational optimization of two long-range interactions first observed in Ac-WINGKWT-NH2, (a) bifurcated H-bonding involving the threonine amide H(N) and side chain OH and the N-terminal acetyl carbonyl and (b) an H-bond between the entgegen-H(N) of the C-terminal amide and the indole ring of Trp6 that stabilizes a face-to-edge indole/indole interaction between Trp1 and Trp6, has afforded < or = 10 residue systems that yield a remarkably stable fold in water. Optimization was achieved by designing a hydrophobic cluster that sequesters these H-bonds from solvent exposure. The structures and extent of amide H/D exchange protection for CH3CH2CO-WI pGXWTGPS (p = D-Pro, X = Leu or Ile) were determined. These two systems are greater than 94% folded at 298 K (97.5% at 280 K) with melting temperatures > 75 degrees C. The fold appears to display minimal fluxionality; a well-converged NMR structure rationalizes all of the large structuring shifts observed, and we suggest that these designed constructs can be viewed as microproteins.
Figures


Similar articles
-
Temperature-dependence of protein hydrogen bond properties as studied by high-resolution NMR.J Mol Biol. 2002 Apr 12;317(5):739-52. doi: 10.1006/jmbi.2002.5446. J Mol Biol. 2002. PMID: 11955021
-
Formation of Annular Protofibrillar Assembly by Cysteine Tripeptide: Unraveling the Interactions with NMR, FTIR, and Molecular Dynamics.J Phys Chem B. 2017 Jul 6;121(26):6367-6379. doi: 10.1021/acs.jpcb.7b04373. Epub 2017 Jun 23. J Phys Chem B. 2017. PMID: 28593765
-
The role of cation-pi interactions in biomolecular association. Design of peptides favoring interactions between cationic and aromatic amino acid side chains.J Am Chem Soc. 2001 Jul 4;123(26):6232-45. doi: 10.1021/ja010401u. J Am Chem Soc. 2001. PMID: 11427046
-
Intramolecular hydrogen bond-controlled prolyl amide isomerization in glucosyl 3'(S)-hydroxy-5'-hydroxymethylproline hybrids: influence of a C-5'-hydroxymethyl substituent on the thermodynamics and kinetics of prolyl amide cis/trans isomerization.J Org Chem. 2009 May 15;74(10):3735-43. doi: 10.1021/jo9003458. J Org Chem. 2009. PMID: 19354261
-
Solid-state NMR and calorimetry of structural waters in helical peptides.J Am Chem Soc. 2002 Mar 13;124(10):2345-51. doi: 10.1021/ja017364r. J Am Chem Soc. 2002. PMID: 11878990
Cited by
-
13C structuring shifts for the analysis of model β-hairpins and β-sheets in proteins: diagnostic shifts appear only at the cross-strand H-bonded residues.J Biomol NMR. 2013 Aug;56(4):313-29. doi: 10.1007/s10858-013-9749-3. Epub 2013 Jul 14. J Biomol NMR. 2013. PMID: 23851979
-
Investigating the role of chain and linker length on the catalytic activity of an H2 production catalyst containing a β-hairpin peptide.J Coord Chem. 2016;69(11-13):1730-1747. doi: 10.1080/00958972.2016.1188924. Epub 2016 Jun 2. J Coord Chem. 2016. PMID: 33093711 Free PMC article.
-
Captides: rigid junctions between beta sheets and small molecules.J Pept Sci. 2014 Sep;20(9):704-15. doi: 10.1002/psc.2657. Epub 2014 Jun 6. J Pept Sci. 2014. PMID: 24909552 Free PMC article.
-
A Conformationally Stable Acyclic β-Hairpin Scaffold Tolerating the Incorporation of Poorly β-Sheet-Prone Amino Acids.Chembiochem. 2022 Feb 16;23(4):e202100604. doi: 10.1002/cbic.202100604. Epub 2021 Dec 16. Chembiochem. 2022. PMID: 34856053 Free PMC article.
-
Circular Permutation of the Trp-cage: Fold Rescue upon Addition of a Hydrophobic Staple.RSC Adv. 2013 Nov 21;2013(43):10.1039/C3RA43674H. doi: 10.1039/C3RA43674H. RSC Adv. 2013. PMID: 24376912 Free PMC article.
References
-
- Kubelka J, Hofrichter J, Eaton WA. Curr Opin Strct Biol. 2004;14:76–88. - PubMed
-
- Struthers MD, Cheng RP, Imperiali B. Science. 1996;271:342–345. - PubMed
- Dahiyat BI, Mayo SL. Science. 1997;278:82–7. - PubMed
- Horng JC, Moroz V, Raleigh DP. J Mol Biol. 2003;326:1261–1270. - PubMed
- Anil B, Craig-Schapiro R, Raleigh DP. J Am Chem Soc. 2006;128:3144–3145. - PubMed
- Jager M, Zhang Y, Bieschke J, Nguyen H, Dendle M, Bowman ME, Noel JP, Gruebele M, Kelly JW. Proc Natl Acad Sci U S A. 2006;103:10648–10653. - PMC - PubMed
- Tang Y, Goger MJ, Raleigh DP. Biochemistry. 2006;45:6940–6946. - PubMed
- Jager M, Dendle M, Fuller AA, Kelly JW. Protein Sci. 2007;16:2306–2313. - PMC - PubMed
- Jager M, Nguyen H, Dendle M, Gruebele M, Kelly JW. Protein Sci. 2007;16:1495–1501. - PMC - PubMed
-
-
The term “miniprotein” has typically been applied to 20 – 45 residue polypeptides that form a stable fold; it has mostly been used for designed systems rather than naturally occurring ones. This term dates back to at least 1996 (Drug Design & Discovery), with a PNAS citation in 1998, and was in common usage prior to our application of it to the Trp-cage fold4. “Microprotein” appears to have two uses in the literature; an older one for specific urinary excretion products and, starting in 2004 (Current Opinions in Biotechnology), as a term for small natural cystine-knot proteins in the plant-cyclotide and conotoxin areas. Herein, following the usual view that “micro” is smaller then “mini”, we use “microprotein” to designated folds with protein-like stability that are much smaller than typical miniproteins.
-
-
- Neidigh JW, Fesinmeyer RM, Andersen NH. Nature Struc Biol. 2002;9:425–430. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

