Probing metagenomics by rapid cluster analysis of very large datasets
- PMID: 18846219
- PMCID: PMC2557142
- DOI: 10.1371/journal.pone.0003375
Probing metagenomics by rapid cluster analysis of very large datasets
Abstract
Background: The scale and diversity of metagenomic sequencing projects challenge both our technical and conceptual approaches in gene and genome annotations. The recent Sorcerer II Global Ocean Sampling (GOS) expedition yielded millions of predicted protein sequences, which significantly altered the landscape of known protein space by more than doubling its size and adding thousands of new families (Yooseph et al., 2007 PLoS Biol 5, e16). Such datasets, not only by their sheer size, but also by many other features, defy conventional analysis and annotation methods.
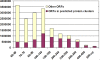
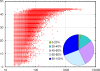
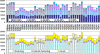
Methodology/principal findings: In this study, we describe an approach for rapid analysis of the sequence diversity and the internal structure of such very large datasets by advanced clustering strategies using the newly modified CD-HIT algorithm. We performed a hierarchical clustering analysis on the 17.4 million Open Reading Frames (ORFs) identified from the GOS study and found over 33 thousand large predicted protein clusters comprising nearly 6 million sequences. Twenty percent of these clusters did not match known protein families by sequence similarity search and might represent novel protein families. Distributions of the large clusters were illustrated on organism composition, functional class, and sample locations.
Conclusion/significance: Our clustering took about two orders of magnitude less computational effort than the similar protein family analysis of original GOS study. This approach will help to analyze other large metagenomic datasets in the future. A Web server with our clustering results and annotations of predicted protein clusters is available online at http://tools.camera.calit2.net/gos under the CAMERA project.
Conflict of interest statement
Figures
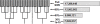


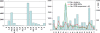
References
-
- DeLong EF, Preston CM, Mincer T, Rich V, Hallam SJ, et al. Community genomics among stratified microbial assemblages in the ocean's interior. Science. 2006;311:496–503. - PubMed

