Transcriptional profiling of mRNA expression in the mouse distal colon
- PMID: 18848557
- PMCID: PMC2748881
- DOI: 10.1053/j.gastro.2008.08.048
Transcriptional profiling of mRNA expression in the mouse distal colon
Abstract
Background & aims: Intestinal epithelial cells and the myenteric plexus of the mouse gastrointestinal tract contain a circadian clock-based intrinsic time-keeping system. Because disruption of the biological clock has been associated with increased susceptibility to colon cancer and gastrointestinal symptoms, we aimed to identify rhythmically expressed genes in the mouse distal colon.
Methods: Microarray analysis was used to identify genes that were rhythmically expressed over a 24-hour light/dark cycle. The transcripts were then classified according to expression pattern, function, and association with physiologic and pathophysiologic processes of the colon.
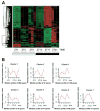
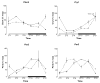
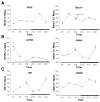
Results: A circadian gene expression pattern was detected in approximately 3.7% of distal colonic genes. A large percentage of these genes were involved in cell signaling, differentiation, and proliferation and cell death. Of all the rhythmically expressed genes in the mouse colon, approximately 7% (64/906) have been associated with colorectal cancer formation (eg, B-cell leukemia/lymphoma-2 [Bcl2]) and 1.8% (18/906) with various colonic functions such as motility and secretion (eg, vasoactive intestinal polypeptide, cystic fibrosis transmembrane conductance regulator).
Conclusions: A subset of genes in the murine colon follows a rhythmic expression pattern. These findings may have significant implications for colonic physiology and pathophysiology.
Figures




Similar articles
-
Clock gene expression in the murine gastrointestinal tract: endogenous rhythmicity and effects of a feeding regimen.Gastroenterology. 2007 Oct;133(4):1250-60. doi: 10.1053/j.gastro.2007.07.009. Epub 2007 Jul 12. Gastroenterology. 2007. PMID: 17919497
-
Insight into the circadian clock within rat colonic epithelial cells.Gastroenterology. 2007 Oct;133(4):1240-9. doi: 10.1053/j.gastro.2007.05.053. Epub 2007 Jun 2. Gastroenterology. 2007. PMID: 17675004
-
Transcriptional Profiling of Daily Patterns of mRNA Expression in the C57BL/6J Mouse Cornea.Curr Eye Res. 2019 Oct;44(10):1054-1066. doi: 10.1080/02713683.2019.1625408. Epub 2019 Jun 25. Curr Eye Res. 2019. PMID: 31136724
-
Role of clock genes in gastrointestinal motility.Am J Physiol Gastrointest Liver Physiol. 2010 Sep;299(3):G549-55. doi: 10.1152/ajpgi.00147.2010. Epub 2010 Jun 17. Am J Physiol Gastrointest Liver Physiol. 2010. PMID: 20558764 Free PMC article. Review.
-
Expression of clock gene products in the suprachiasmatic nucleus in relation to circadian behaviour.Novartis Found Symp. 2003;253:203-17; discussion 102-9, 218-22, 281-4. doi: 10.1002/0470090839.ch15. Novartis Found Symp. 2003. PMID: 14712923 Review.
Cited by
-
Integration of photoperiod and time-restricted feeding on the circadian gene rhythms in juvenile salmon.Sci Rep. 2025 May 9;15(1):16156. doi: 10.1038/s41598-025-01069-0. Sci Rep. 2025. PMID: 40346079 Free PMC article.
-
Nocturnin regulates circadian trafficking of dietary lipid in intestinal enterocytes.Curr Biol. 2011 Aug 23;21(16):1347-55. doi: 10.1016/j.cub.2011.07.018. Epub 2011 Aug 4. Curr Biol. 2011. PMID: 21820310 Free PMC article.
-
The role of cryptochrome (CRY) in cancer: molecular mechanisms and clock-based therapeutic strategies.Acta Biochim Biophys Sin (Shanghai). 2025 Mar 19;57(7):1037-1046. doi: 10.3724/abbs.2025025. Acta Biochim Biophys Sin (Shanghai). 2025. PMID: 40109093 Free PMC article. Review.
-
Importance of Circadian Rhythms in the Ocular Surface.Biomolecules. 2024 Jul 4;14(7):796. doi: 10.3390/biom14070796. Biomolecules. 2024. PMID: 39062510 Free PMC article. Review.
-
BMAL1 Regulates the Daily Timing of Colitis.Front Cell Infect Microbiol. 2022 Feb 9;12:773413. doi: 10.3389/fcimb.2022.773413. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 35223537 Free PMC article.
References
-
- Hoogerwerf WA. Biologic clocks and the gut. Curr Gastroenterol Rep. 2006;8:353–359. - PubMed
-
- Scheving LA. Biological clocks and the digestive system. Gastroenterology. 2000;119:536–549. - PubMed
-
- Ko CH, Takahashi JS. Molecular components of the mammalian circadian clock. Hum Mol Genet. 2006;15:R271–R277. - PubMed
-
- Albrecht U, Eichele G. The mammalian circadian clock. Curr Opin Genet Dev. 2003;13:271–277. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

