Structural dynamics of the ribosome
- PMID: 18848900
- PMCID: PMC3923522
- DOI: 10.1016/j.cbpa.2008.08.037
Structural dynamics of the ribosome
Abstract
Protein synthesis is inherently a dynamic process, requiring both small-scale and large-scale movements of tRNA and mRNA. It has long been suspected that these movements might be coupled to conformational changes in the ribosome, and in its RNA moieties in particular. Recently, the nature of ribosome structural dynamics has begun to emerge from a combination of approaches, most notably cryo-EM, X-ray crystallography, and FRET. Ribosome movement occurs both on a grand scale, as in the intersubunit rotational movements that are coupled to tRNA-mRNA translocation, and in intricate localized rearrangements such as those that accompany codon-anticodon recognition and peptide bond formation. In spite of much progress, our understanding of the mechanics of translation is now beset with countless new questions, reflecting the vast molecular architecture of the ribosome itself.
Figures


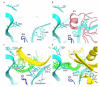

References
-
- Green R, Noller HF. Ribosomes and translation. Annu Rev Biochem. 1997;66:679–716. - PubMed
-
- Ogle JM, Carter AP, Ramakrishnan V. Insights into the decoding mechanism from recent ribosome structures. Trends Biochem Sci. 2003;28:259–266. - PubMed
-
- Ban N, Nissen P, Hansen J, Moore PB, Steitz TA. The complete atomic structure of the large ribosomal subunit at 2.4 A resolution. Science. 2000;289:905–920. - PubMed
-
- Yarus M. Boundaries for an RNA world. Curr Opin Chem Biol. 1999;3:260–267. - PubMed
-
- Bretscher MS. Translocation in protein synthesis: a hybrid structure model. Nature. 1968;218:675–677. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

