A genome-wide screen for beta-catenin binding sites identifies a downstream enhancer element that controls c-Myc gene expression
- PMID: 18852287
- PMCID: PMC2593444
- DOI: 10.1128/MCB.00744-08
A genome-wide screen for beta-catenin binding sites identifies a downstream enhancer element that controls c-Myc gene expression
Abstract
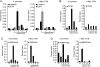
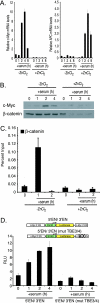
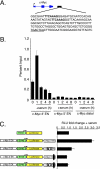
Mutations in components of the Wnt signaling pathway initiate colorectal carcinogenesis by deregulating the beta-catenin transcriptional coactivator. beta-Catenin activation of one target in particular, the c-Myc proto-oncogene, is required for colon cancer pathogenesis. beta-Catenin is known to regulate c-Myc expression via sequences upstream of the transcription start site. Here, we report that a more robust beta-catenin binding region localizes 1.4 kb downstream from the c-Myc transcriptional stop site. This site was discovered using a genome-wide method for identifying transcription factor binding sites termed serial analysis of chromatin occupancy. Chromatin immunoprecipitation-scanning assays demonstrate that the 5' enhancer and the 3' binding element are the only beta-catenin and TCF4 binding regions across the c-Myc locus. When placed downstream of a simian virus 40-driven promoter-luciferase construct, the 3' element activated luciferase transcription when introduced into HCT116 cells. c-Myc transcription is negligible in quiescent HCT116 cells but is induced when cells reenter the cell cycle after the addition of mitogens. Using these cells, we found that beta-catenin and TCF4 occupancy at the 3' enhancer precede occupancy at the 5' enhancer. Association of c-Jun, beta-catenin, and TCF4 specifically with the downstream enhancer underlies mitogen stimulation of c-Myc transcription. Our findings indicate that a downstream enhancer element provides the principal regulation of c-Myc expression.
Figures





Similar articles
-
A dynamic exchange of TCF3 and TCF4 transcription factors controls MYC expression in colorectal cancer cells.Cell Cycle. 2015;14(3):323-32. doi: 10.4161/15384101.2014.980643. Cell Cycle. 2015. PMID: 25659031 Free PMC article.
-
A beta-catenin/TCF-coordinated chromatin loop at MYC integrates 5' and 3' Wnt responsive enhancers.Proc Natl Acad Sci U S A. 2010 Jan 5;107(1):145-50. doi: 10.1073/pnas.0912294107. Epub 2009 Dec 4. Proc Natl Acad Sci U S A. 2010. PMID: 19966299 Free PMC article.
-
VDR/RXR and TCF4/β-catenin cistromes in colonic cells of colorectal tumor origin: impact on c-FOS and c-MYC gene expression.Mol Endocrinol. 2012 Jan;26(1):37-51. doi: 10.1210/me.2011-1109. Epub 2011 Nov 22. Mol Endocrinol. 2012. PMID: 22108803 Free PMC article.
-
SPDEF Induces Quiescence of Colorectal Cancer Cells by Changing the Transcriptional Targets of β-catenin.Gastroenterology. 2017 Jul;153(1):205-218.e8. doi: 10.1053/j.gastro.2017.03.048. Epub 2017 Apr 5. Gastroenterology. 2017. PMID: 28390865 Free PMC article.
-
The links between transcription, beta-catenin/JNK signaling, and carcinogenesis.Mol Cancer Res. 2009 Aug;7(8):1189-96. doi: 10.1158/1541-7786.MCR-09-0027. Epub 2009 Aug 11. Mol Cancer Res. 2009. PMID: 19671687 Review.
Cited by
-
WNT/β-Catenin Signaling Regulates Multiple Steps of Myogenesis by Regulating Step-Specific Targets.Mol Cell Biol. 2015 May;35(10):1763-76. doi: 10.1128/MCB.01180-14. Epub 2015 Mar 9. Mol Cell Biol. 2015. PMID: 25755281 Free PMC article.
-
The antiproliferative effect of indomethacin-loaded lipid-core nanocapsules in glioma cells is mediated by cell cycle regulation, differentiation, and the inhibition of survival pathways.Int J Nanomedicine. 2013;8:711-28. doi: 10.2147/IJN.S40284. Epub 2013 Feb 18. Int J Nanomedicine. 2013. PMID: 23440594 Free PMC article.
-
Inferring regulatory element landscapes and transcription factor networks from cancer methylomes.Genome Biol. 2015 May 21;16(1):105. doi: 10.1186/s13059-015-0668-3. Genome Biol. 2015. PMID: 25994056 Free PMC article.
-
DLK1/DIO3 locus upregulation by a β-catenin-dependent enhancer drives cell proliferation and liver tumorigenesis.Mol Ther. 2024 Apr 3;32(4):1125-1143. doi: 10.1016/j.ymthe.2024.01.036. Epub 2024 Feb 3. Mol Ther. 2024. PMID: 38311851 Free PMC article.
-
The Crohn's disease associated SNP rs6651252 impacts MYC gene expression in human colonic epithelial cells.PLoS One. 2019 Feb 22;14(2):e0212850. doi: 10.1371/journal.pone.0212850. eCollection 2019. PLoS One. 2019. PMID: 30794691 Free PMC article.
References
-
- Armstrong, B. C., and G. W. Krystal. 1992. Isolation and characterization of complementary DNA for N-cym, a gene encoded by the DNA strand opposite to N-myc. Cell Growth Differ. 3385-390. - PubMed
-
- Banerji, J., S. Rusconi, and W. Schaffner. 1981. Expression of a beta-globin gene is enhanced by remote SV40 DNA sequences. Cell 27299-308. - PubMed
-
- Barski, A., S. Cuddapah, K. Cui, T. Y. Roh, D. E. Schones, Z. Wang, G. Wei, I. Chepelev, and K. Zhao. 2007. High-resolution profiling of histone methylations in the human genome. Cell 129823-837. - PubMed
-
- Berger, S. L. 2007. The complex language of chromatin regulation during transcription. Nature 447407-412. - PubMed
-
- Bommer, G. T., and E. R. Fearon. 2007. Role of c-Myc in Apc mutant intestinal phenotype: case closed or time for a new beginning? Cancer Cell 11391-394. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
