Rapid multiplex PCR and real-time TaqMan PCR assays for detection of Salmonella enterica and the highly virulent serovars Choleraesuis and Paratyphi C
- PMID: 18923008
- PMCID: PMC2593286
- DOI: 10.1128/JCM.01229-08
Rapid multiplex PCR and real-time TaqMan PCR assays for detection of Salmonella enterica and the highly virulent serovars Choleraesuis and Paratyphi C
Abstract
Salmonella enterica is a human pathogen with over 2,500 serovars characterized. S. enterica serovars Choleraesuis and Paratyphi C are two globally distributed serovars. We have developed a rapid molecular-typing method to detect serovars Choleraesuis and Paratyphi C in food samples by using a comparative-genomics approach to identify regions unique to each serovar from the sequenced genomes. A Salmonella-specific primer pair based on oriC was designed as an internal control to establish accuracy, sensitivity, and reproducibility. Serovar-specific primer sets based on regions of difference between serovars Choleraesuis and Paratyphi C were designed for real-time PCR assays. Three primer sets were used to screen a collection of over 100 Salmonella strains, and both serovars Choleraesuis and Paratyphi C gave unique amplification patterns. To develop the technique for practical use, its sensitivity for detection of Salmonella spp. in a food matrix was determined by spiking experiments. The technique was also adapted for a real-time PCR rapid-detection assay for both serovars Choleraesuis and Paratyphi C that complements the current procedures for Salmonella sp. isolation and serotyping.
Figures

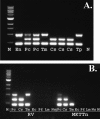

References
-
- Boyd, E. F., F. S. Wang, P. Beltran, S. A. Plock, K. Nelson, and R. K. Selander. 1993. Salmonella reference collection B (SARB): strains of 37 serovars of subspecies I. J. Gen. Microbiol. 1391125-1132. - PubMed
-
- Centers For Disease Control and Prevention. 2008. Multistate outbreak of human Salmonella infections caused by contaminated dry dog food-United States, 2006-2007. MMWR Morb. Mortal. Wkly. Rep. 57521-524. - PubMed
-
- Chen, P. L., C. J. Wu, C. M. Chang, H. C. Lee, N. Y. Lee, H. I. Shih, C. C. Lee, N. Y. Ko, L. R. Wang, and W. C. Ko. 2007. Extraintestinal focal infections in adults with Salmonella enterica serotype Choleraesuis bacteremia. J. Microbiol. Immunol. Infect. 40240-247. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

