Long-range enhancer associated with chromatin looping allows AP-1 regulation of the peptidylarginine deiminase 3 gene in differentiated keratinocyte
- PMID: 18923650
- PMCID: PMC2566589
- DOI: 10.1371/journal.pone.0003408
Long-range enhancer associated with chromatin looping allows AP-1 regulation of the peptidylarginine deiminase 3 gene in differentiated keratinocyte
Abstract
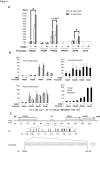
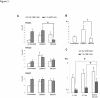
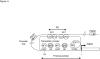
Transcription control at a distance is a critical mechanism, particularly for contiguous genes. The peptidylarginine deiminases (PADs) catalyse the conversion of protein-bound arginine into citrulline (deimination), a critical reaction in the pathophysiology of multiple sclerosis, Alzheimer's disease and rheumatoid arthritis, and in the metabolism of the major epidermal barrier protein filaggrin, a strong predisposing factor for atopic dermatitis. PADs are encoded by 5 clustered PADI genes (1p35-6). Unclear are the mechanisms controlling the expression of the gene PADI3 encoding the PAD3 isoform, a strong candidate for the deimination of filaggrin in the terminally differentiating epidermal keratinocyte. We describe the first PAD Intergenic Enhancer (PIE), an evolutionary conserved non coding segment located 86-kb from the PADI3 promoter. PIE is a strong enhancer of the PADI3 promoter in Ca2+-differentiated epidermal keratinocytes, and requires bound AP-1 factors, namely c-Jun and c-Fos. As compared to proliferative keratinocytes, calcium stimulation specifically associates with increased local DNase I hypersensitivity around PIE, and increased physical proximity of PIE and PADI3 as assessed by Chromosome Conformation Capture. The specific AP-1 inhibitor nordihydroguaiaretic acid suppresses the calcium-induced increase of PADI3 mRNA levels in keratinocytes. Our findings pave the way to the exploration of deimination control during tumorigenesis and wound healing, two conditions for which AP-1 factors are critical, and disclose that long-range transcription control has a role in the regulation of the gene PADI3. Since invalidation of distant regulators causes a variety of human diseases, PIE results to be a plausible candidate in association studies on deimination-related disorders or atopic disease.
Conflict of interest statement
Figures



References
-
- Chavanas S, Mechin MC, Takahara H, Kawada A, Nachat R, et al. Comparative analysis of the mouse and human peptidylarginine deiminase gene clusters reveals highly conserved non-coding segments and a new human gene, PADI6. Gene. 2004;330:19–27. - PubMed
-
- Liu GY, Liao YF, Chang WH, Liu CC, Hsieh MC, et al. Overexpression of peptidylarginine deiminase IV features in apoptosis of haematopoietic cells. Apoptosis. 2006;11:183–196. - PubMed
-
- Cuthbert GL, Daujat S, Snowden AW, Erdjument-Bromage H, Hagiwara T, et al. Histone deimination antagonizes arginine methylation. Cell. 2004;118:545–553. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

