Gene expression profiling of two distinct neuronal populations in the rodent spinal cord
- PMID: 18923679
- PMCID: PMC2566599
- DOI: 10.1371/journal.pone.0003415
Gene expression profiling of two distinct neuronal populations in the rodent spinal cord
Abstract
Background: In the field of neuroscience microarray gene expression profiles on anatomically defined brain structures are being used increasingly to study both normal brain functions as well as pathological states. Fluorescent tracing techniques in brain tissue that identifies distinct neuronal populations can in combination with global gene expression profiling potentially increase the resolution and specificity of such studies to shed new light on neuronal functions at the cellular level.
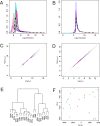
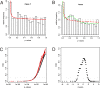
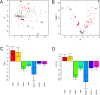
Methodology/principal findings: We examine the microarray gene expression profiles of two distinct neuronal populations in the spinal cord of the neonatal rat, the principal motor neurons and specific interneurons involved in motor control. The gene expression profiles of the respective cell populations were obtained from amplified mRNA originating from 50-250 fluorescently identified and laser microdissected cells. In the data analysis we combine a new microarray normalization procedure with a conglomerate measure of significant differential gene expression. Using our methodology we find 32 genes to be more expressed in the interneurons compared to the motor neurons that all except one have not previously been associated with this neuronal population. As a validation of our method we find 17 genes to be more expressed in the motor neurons than in the interneurons and of these only one had not previously been described in this population.
Conclusions/significance: We provide an optimized experimental protocol that allows isolation of gene transcripts from fluorescent retrogradely labeled cell populations in fresh tissue, which can be used to generate amplified aRNA for microarray hybridization from as few as 50 laser microdissected cells. Using this optimized experimental protocol in combination with our microarray analysis methodology we find 49 differentially expressed genes between the motor neurons and the interneurons that reflect the functional differences between these two cell populations in generating and transmitting the motor output in the rodent spinal cord.
Conflict of interest statement
Figures

References
-
- Diaz E. A functional genomics guide to the galaxy of neuronal cell types. Nat Neurosci. 2006;9:10–12. - PubMed
-
- Shoemaker DD, Linsley PS. Recent developments in DNA microarrays. Curr Opin Microbiol. 2002;5:334–337. - PubMed
-
- Nelson SB, Hempel C, Sugino K. Probing the transcriptome of neuronal cell types. Curr Opin Neurobiol. 2006;16:571–576. - PubMed
-
- Nelson SB, Sugino K, Hempel CM. The problem of neuronal cell types: a physiological genomics approach. Trends Neurosci. 2006;29:339–345. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

