Does intra-individual major histocompatibility complex diversity keep a golden mean?
- PMID: 18926972
- PMCID: PMC2666699
- DOI: 10.1098/rstb.2008.0174
Does intra-individual major histocompatibility complex diversity keep a golden mean?
Abstract
An adaptive immune response is usually initiated only if a major histocompatibility complex (MHC) molecule presents pathogen-derived peptides to T-cells. Every MHC molecule can present only peptides that match its peptide-binding groove. Thus, it seems advantageous for an individual to express many different MHC molecules to be able to resist many different pathogens. However, although MHC genes are the most polymorphic genes of vertebrates, each individual has only a very small subset of the diversity at the population level. This is an evolutionary paradox. We provide an overview of the current data on infection studies and mate-choice experiments and conclude that overall evidence suggests that intermediate intra-individual MHC diversity is optimal. Selective forces that may set an upper limit to intra-individual MHC diversity are discussed. An updated mathematical model based on recent findings on T-cell selection can predict the natural range of intra-individual MHC diversity. Thus, the aim of our review is to evaluate whether the number of MHC alleles usually present in individuals may be optimal to balance the advantages of presenting an increased range of peptides versus the disadvantages of an increased loss of T-cells.
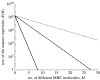
Figures


Similar articles
-
Evolution of major histocompatibility complex gene copy number.PLoS Comput Biol. 2019 May 16;15(5):e1007015. doi: 10.1371/journal.pcbi.1007015. eCollection 2019 May. PLoS Comput Biol. 2019. PMID: 31095555 Free PMC article.
-
Antigen presentation on MHC molecules as a diversity filter that enhances immune efficacy.J Theor Biol. 2003 Sep 21;224(2):249-67. doi: 10.1016/s0022-5193(03)00162-0. J Theor Biol. 2003. PMID: 12927531
-
Pathogen burden, co-infection and major histocompatibility complex variability in the European badger (Meles meles).Mol Ecol. 2014 Oct;23(20):5072-88. doi: 10.1111/mec.12917. Epub 2014 Oct 7. Mol Ecol. 2014. PMID: 25211523
-
The Nature of Selection on the Major Histocompatibility Complex.Crit Rev Immunol. 2017;37(2-6):75-120. doi: 10.1615/CritRevImmunol.v37.i2-6.10. Crit Rev Immunol. 2017. PMID: 29773018 Review.
-
Natural selection and the diversification of vertebrate immune effectors.Immunol Rev. 2002 Dec;190:161-8. doi: 10.1034/j.1600-065x.2002.19012.x. Immunol Rev. 2002. PMID: 12493013 Review.
Cited by
-
Evolutionary trade-offs constraining the MHC gene expansion: beyond simple TCR depletion model.Front Immunol. 2024 Jan 8;14:1240723. doi: 10.3389/fimmu.2023.1240723. eCollection 2023. Front Immunol. 2024. PMID: 38259496 Free PMC article.
-
Divergent allele advantage at MHC-DRB through direct and maternal genotypic effects and its consequences for allele pool composition and mating.Proc Biol Sci. 2013 Jul 7;280(1762):20130714. doi: 10.1098/rspb.2013.0714. Print 2013 Jul 7. Proc Biol Sci. 2013. PMID: 23677346 Free PMC article.
-
Cryptic haplotype-specific gamete selection yields offspring with optimal MHC immune genes.Evolution. 2018 Nov;72(11):2478-2490. doi: 10.1111/evo.13591. Epub 2018 Sep 24. Evolution. 2018. PMID: 30246285 Free PMC article.
-
Mhc supertypes confer both qualitative and quantitative resistance to avian malaria infections in a wild bird population.Proc Biol Sci. 2013 Mar 20;280(1759):20130134. doi: 10.1098/rspb.2013.0134. Print 2013 May 22. Proc Biol Sci. 2013. PMID: 23516242 Free PMC article.
-
Evolution of Copy Number at the MHC Varies across the Avian Tree of Life.Genome Biol Evol. 2019 Jan 1;11(1):17-28. doi: 10.1093/gbe/evy253. Genome Biol Evol. 2019. PMID: 30476037 Free PMC article.
References
-
- Aeschlimann P.B., Häberli M.A., Reusch T.B.H., Boehm T., Milinski M. Female sticklebacks Gasterosteus aculeatus use self-reference to optimize MHC allele number during mate selection. Behav. Ecol. Sociobiol. 2003;54:119–126. doi:10.1007/s00265-003-0611-6 - DOI
-
- Allen T.M., et al. Tat-specific cytotoxic T lymphocytes select for SIV escape variants during resolution of primary viraemia. Nature. 2000;407:386–390. doi:10.1038/35036559 - DOI - PubMed
-
- Aoyagi K., Dijkstra J.M., Xia C., Denda I., Ototake M., Hashimoto K., Nakanishi T. Classical MHC class I genes composed of highly divergent sequence lineages share a single locus in rainbow trout (Oncorhynchus mykiss) J. Immunol. 2002;168:260–273. - PubMed
-
- Blattman J.N., Antia R., Sourdive D.J.D., Wang X.C., Kaech S.M., Murali-Krishna K., Altman J.D., Ahmed R. Estimating the precursor frequency of naive antigen-specific CD8 T cells. J. Exp. Med. 2002;195:657–664. doi:10.1084/jem.20001021 - DOI - PMC - PubMed
-
- Bonneaud C., Mazuc J., Chastel O., Westerdahl H., Sorci G. Terminal investment induced by immune challenge and fitness traits associated with major histocompatibility complex in the house sparrow. Evolution. 2004;58:2823–2830. doi:10.1554/04-279 - DOI - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials

