In vivo cloning of artificial DNA nanostructures
- PMID: 18927233
- PMCID: PMC2584761
- DOI: 10.1073/pnas.0805416105
In vivo cloning of artificial DNA nanostructures
Abstract
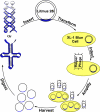
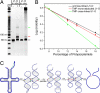
Mimicking nature is both a key goal and a difficult challenge for the scientific enterprise. DNA, well known as the genetic-information carrier in nature, can be replicated efficiently in living cells. Today, despite the dramatic evolution of DNA nanotechnology, a versatile method that replicates artificial DNA nanostructures with complex secondary structures remains an appealing target. Previous success in replicating DNA nanostructures enzymatically in vitro suggests that a possible solution could be cloning these nanostructures by using viruses. Here, we report a system where a single-stranded DNA nanostructure (Holliday junction or paranemic cross-over DNA) is inserted into a phagemid, transformed into XL1-Blue cells and amplified in vivo in the presence of helper phages. High copy numbers of cloned nanostructures can be obtained readily by using standard molecular biology techniques. Correct replication is verified by a number of assays including nondenaturing PAGE, Ferguson analysis, endonuclease VII digestion, and hydroxyl radical autofootprinting. The simplicity, efficiency, and fidelity of nature are fully reflected in this system. UV-induced psoralen cross-linking is used to probe the secondary structure of the inserted junction in infected cells. Our data suggest the possible formation of the immobile four-arm junction in vivo.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Comment in
-
Rolling out DNA nanostructures in vivo.Proc Natl Acad Sci U S A. 2008 Nov 18;105(46):17593-4. doi: 10.1073/pnas.0810029105. Epub 2008 Nov 13. Proc Natl Acad Sci U S A. 2008. PMID: 19008359 Free PMC article. No abstract available.
References
-
- Seeman NC. DNA in a material world. Nature. 2003;421:427–431. - PubMed
-
- Lin C, Liu Y, Rinker S, Yan H. DNA tile based self-assembly: Building complex nanoarchitectures. ChemPhysChem. 2006;7:1641–1647. - PubMed
-
- Chen J, Seeman NC. The synthesis from DNA of a molecule with the connectivity of a cube. Nature. 1991;350:631–633. - PubMed
-
- Rothemund PWK. Folding DNA to create nanoscale shapes and patterns. Nature. 2006;440:297–302. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

