Complex regulatory pathways coordinate cell-cycle progression and development in Caulobacter crescentus
- PMID: 18929067
- PMCID: PMC2621326
- DOI: 10.1016/S0065-2911(08)00001-5
Complex regulatory pathways coordinate cell-cycle progression and development in Caulobacter crescentus
Abstract
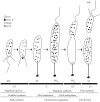
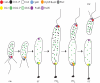
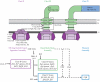
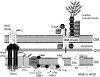
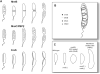
Caulobacter crescentus has become the predominant bacterial model system to study the regulation of cell-cycle progression. Stage-specific processes such as chromosome replication and segregation, and cell division are coordinated with the development of four polar structures: the flagellum, pili, stalk, and holdfast. The production, activation, localization, and proteolysis of specific regulatory proteins at precise times during the cell cycle culminate in the ability of the cell to produce two physiologically distinct daughter cells. We examine the recent advances that have enhanced our understanding of the mechanisms of temporal and spatial regulation that occur during cell-cycle progression.
Figures






References
-
- Aaron M, Charbon G, Lam H, Schwarz H, Vollmer W, Jacobs-Wagner C. The tubulin homologue FtsZ contributes to cell elongation by guiding cell wall precursor synthesis in Caulobacter crescentus. Mol. Microbiol. 2007;64:938–952. - PubMed
-
- Abraham WR, Strompl C, Meyer H, Lindholst S, Moore ER, Christ R, Vancanneyt M, Tindall BJ, Bennasar A, Smit J, Tesar M. Phylogeny and polyphasic taxonomy of Caulobacter species. Proposal of Maricaulis gen. nov. with Maricaulis maris (Poindexter) comb. nov. as the type species, and emended description of the genera Brevundimonas and Caulobacter. Int. J. Syst. Bacteriol. 1999;49(Pt 3):1053–73. - PubMed
-
- Addinall SG, Holland B. The tubulin ancester, FtsZ, draughtsman, designer and driving force for bacterial cytokinesis. J. Mol. Biol. 2002;318:219–236. - PubMed
-
- Aldridge P, Hughes KT. Regulation of flagellar assembly. Currents Opinion in Microbiology. 2002;5:160–5. - PubMed
-
- Aldridge P, Jenal U. Cell cycle-dependent degradation of a flagellar motor component requires a novel-type response regulator. Mol. Microbiol. 1999;32:379–392. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

