Evaluation of strong cation exchange versus isoelectric focusing of peptides for multidimensional liquid chromatography-tandem mass spectrometry
- PMID: 18939861
- PMCID: PMC2669493
- DOI: 10.1021/pr8004666
Evaluation of strong cation exchange versus isoelectric focusing of peptides for multidimensional liquid chromatography-tandem mass spectrometry
Abstract
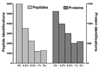
Shotgun proteome analysis platforms based on multidimensional liquid chromatography-tandem mass spectrometry (LC-MS/MS) provide a powerful means to discover biomarker candidates in tissue specimens. Analysis platforms must balance sensitivity for peptide detection, reproducibility of detected peptide inventories and analytical throughput for protein amounts commonly present in tissue biospecimens (< 100 microg), such that platform stability is sufficient to detect modest changes in complex proteomes. We compared shotgun proteomics platforms by analyzing tryptic digests of whole cell and tissue proteomes using strong cation exchange (SCX) and isoelectric focusing (IEF) separations of peptides prior to LC-MS/MS analysis on a LTQ-Orbitrap hybrid instrument. IEF separations provided superior reproducibility and resolution for peptide fractionation from samples corresponding to both large (100 microg) and small (10 microg) protein inputs. SCX generated more peptide and protein identifications than did IEF with small (10 microg) samples, whereas the two platforms yielded similar numbers of identifications with large (100 microg) samples. In nine replicate analyses of tryptic peptides from 50 microg colon adenocarcinoma protein, overlap in protein detection by the two platforms was 77% of all proteins detected by both methods combined. IEF more quickly approached maximal detection, with 90% of IEF-detectable medium abundance proteins (those detected with a total of 3-4 peptides) detected within three replicate analyses. In contrast, the SCX platform required six replicates to detect 90% of SCX-detectable medium abundance proteins. High reproducibility and efficient resolution of IEF peptide separations make the IEF platform superior to the SCX platform for biomarker discovery via shotgun proteomic analyses of tissue specimens.
Figures





References
-
- Rifai N, Gillette MA, Carr SA. Protein biomarker discovery and validation: the long and uncertain path to clinical utility. Nat Biotechnol. 2006;24(8):971–983. - PubMed
-
- Cravatt BF, Simon GM, Yates JR., 3rd The biological impact of mass-spectrometry-based proteomics. Nature. 2007;450(7172):991–1000. - PubMed
-
- Omenn GS, States DJ, Adamski M, Blackwell TW, Menon R, Hermjakob H, Apweiler R, Haab BB, Simpson RJ, Eddes JS, Kapp EA, Moritz RL, Chan DW, Rai AJ, Admon A, Aebersold R, Eng J, Hancock WS, Hefta SA, Meyer H, Paik YK, Yoo JS, Ping P, Pounds J, Adkins J, Qian X, Wang R, Wasinger V, Wu CY, Zhao X, Zeng R, Archakov A, Tsugita A, Beer I, Pandey A, Pisano M, Andrews P, Tammen H, Speicher DW, Hanash SM. Overview of the HUPO Plasma Proteome Project: results from the pilot phase with 35 collaborating laboratories and multiple analytical groups, generating a core dataset of 3020 proteins and a publicly-available database. Proteomics. 2005;5(13):3226–3245. - PubMed
-
- Taylor SW, Fahy E, Zhang B, Glenn GM, Warnock DE, Wiley S, Murphy AN, Gaucher SP, Capaldi RA, Gibson BW, Ghosh SS. Characterization of the human heart mitochondrial proteome. Nat Biotechnol. 2003;21(3):281–286. - PubMed
-
- Hall N, Karras M, Raine JD, Carlton JM, Kooij TW, Berriman M, Florens L, Janssen CS, Pain A, Christophides GK, James K, Rutherford K, Harris B, Harris D, Churcher C, Quail MA, Ormond D, Doggett J, Trueman HE, Mendoza J, Bidwell SL, Rajandream MA, Carucci DJ, Yates JR, 3rd, Kafatos FC, Janse CJ, Barrell B, Turner CM, Waters AP, Sinden RE. A comprehensive survey of the Plasmodium life cycle by genomic, transcriptomic, and proteomic analyses. Science. 2005;307(5706):82–86. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

