A distinct translation initiation mechanism generates cryptic peptides for immune surveillance
- PMID: 18941630
- PMCID: PMC2565129
- DOI: 10.1371/journal.pone.0003460
A distinct translation initiation mechanism generates cryptic peptides for immune surveillance
Abstract
MHC class I molecules present a comprehensive mixture of peptides on the cell surface for immune surveillance. The peptides represent the intracellular protein milieu produced by translation of endogenous mRNAs. Unexpectedly, the peptides are encoded not only in conventional AUG initiated translational reading frames but also in alternative cryptic reading frames. Here, we analyzed how ribosomes recognize and use cryptic initiation codons in the mRNA. We find that translation initiation complexes assemble at non-AUG codons but differ from canonical AUG initiation in response to specific inhibitors acting within the peptidyl transferase and decoding centers of the ribosome. Thus, cryptic translation at non-AUG start codons can utilize a distinct initiation mechanism which could be differentially regulated to provide peptides for immune surveillance.
Conflict of interest statement
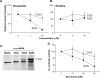
Figures





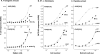
Similar articles
-
Unanticipated antigens: translation initiation at CUG with leucine.PLoS Biol. 2004 Nov;2(11):e366. doi: 10.1371/journal.pbio.0020366. Epub 2004 Oct 26. PLoS Biol. 2004. PMID: 15510226 Free PMC article.
-
Leucine-tRNA initiates at CUG start codons for protein synthesis and presentation by MHC class I.Science. 2012 Jun 29;336(6089):1719-23. doi: 10.1126/science.1220270. Science. 2012. PMID: 22745432
-
Presentation of out-of-frame peptide/MHC class I complexes by a novel translation initiation mechanism.Immunity. 1999 Jun;10(6):681-90. doi: 10.1016/s1074-7613(00)80067-9. Immunity. 1999. PMID: 10403643
-
Non-AUG translation initiation in mammals.Genome Biol. 2022 May 9;23(1):111. doi: 10.1186/s13059-022-02674-2. Genome Biol. 2022. PMID: 35534899 Free PMC article. Review.
-
Translation initiation at AUG and non-AUG triplets in plants.Plant Sci. 2023 Oct;335:111822. doi: 10.1016/j.plantsci.2023.111822. Epub 2023 Aug 14. Plant Sci. 2023. PMID: 37574140 Review.
Cited by
-
Comprehensive profiling of translation initiation in influenza virus infected cells.PLoS Pathog. 2019 Jan 23;15(1):e1007518. doi: 10.1371/journal.ppat.1007518. eCollection 2019 Jan. PLoS Pathog. 2019. PMID: 30673779 Free PMC article.
-
Antisense-Derived HIV-1 Cryptic Epitopes Are Not Major Drivers of Viral Evolution during the Acute Phase of Infection.J Virol. 2018 Sep 12;92(19):e00711-18. doi: 10.1128/JVI.00711-18. Print 2018 Oct 1. J Virol. 2018. PMID: 30021907 Free PMC article.
-
Crystal structure of the eukaryotic translation initiation factor 2A from Schizosaccharomyces pombe.J Struct Funct Genomics. 2014 Sep;15(3):125-30. doi: 10.1007/s10969-014-9177-y. Epub 2014 Feb 26. J Struct Funct Genomics. 2014. PMID: 24569939 Free PMC article.
-
RAN translation and frameshifting as translational challenges at simple repeats of human neurodegenerative disorders.Nucleic Acids Res. 2014 Oct 29;42(19):11849-64. doi: 10.1093/nar/gku794. Epub 2014 Sep 12. Nucleic Acids Res. 2014. PMID: 25217582 Free PMC article. Review.
-
Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes.Cell. 2011 Nov 11;147(4):789-802. doi: 10.1016/j.cell.2011.10.002. Epub 2011 Nov 3. Cell. 2011. PMID: 22056041 Free PMC article.
References
-
- Shastri N, Schwab S, Serwold T. Producing nature's gene-chips. The generation of peptides for display by MHC class I molecules. Annu Rev Immunol. 2002;20:463–493. - PubMed
-
- Cresswell P, Ackerman AL, Giodini A, Peaper DR, Wearsch PA. Mechanisms of MHC class I-restricted antigen processing and cross-presentation. Immunol Rev. 2005;207:145–157. - PubMed
-
- Jensen PE. Recent advances in antigen processing and presentation. Nat Immunol. 2007;8:1041–1048. - PubMed
-
- Yewdell JW, Reits E, Neefjes J. Making sense of mass destruction: quantitating MHC class I antigen presentation. Nat Rev Immunol. 2003;3:952–961. - PubMed
-
- Rock KL, York IA, Saric T, Goldberg AL. Protein degradation and the generation of MHC class I-presented peptides. Adv Immunol. 2002;80:1–70. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

