Mutation of H63 and its catalytic affect on the methionine aminopeptidase from Escherichia coli
- PMID: 18952013
- PMCID: PMC2674292
- DOI: 10.1016/j.bbapap.2008.09.012
Mutation of H63 and its catalytic affect on the methionine aminopeptidase from Escherichia coli
Abstract
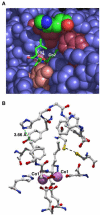
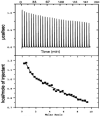
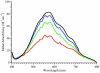
In order to gain insight into the mechanistic role of a flexible exterior loop near the active site, made up of Y62, H63, G64, and Y65, that has been proposed to play an important role in substrate binding and recognition in the methionyl aminopeptidase from Escherichia coli (EcMetAP-I), the H63A enzyme was prepared. Mutation of H63 to alanine does not affect the ability of the enzyme to bind divalent metal ions. The specific activity of H63A EcMetAP-I was determined using four different substrates of varying lengths, namely, l-Met-p-NA, MAS, MGMM and MSSHRWDW. For the smallest/shortest substrate (l-Met-p-NA) the specific activity decreased nearly seven fold but as the peptide length increased, the specific activity also increased and became comparable to WT EcMetAP-I. This decrease in specific activity is primarily due to a decrease in the observed k(cat) values, which decreases nearly sixty-fold for l-Met-p-NA while only a four-fold decrease is observed for the tri- and tetra-peptide substrates. Interestingly, no change in k(cat) was observed when the octa-peptide MSSHRWDW was used as a substrate. These data suggest that H63 affects the hydrolysis of small peptide substrates whereas large peptides can overcome the observed loss in binding energy, as predicted from K(m) values, by additional hydrophilic and hydrophobic interactions.
Figures


Similar articles
-
Kinetic and spectroscopic analysis of the catalytic role of H79 in the methionine aminopeptidase from Escherichia coli.Biochemistry. 2008 Nov 11;47(45):11885-93. doi: 10.1021/bi801499g. Epub 2008 Oct 15. Biochemistry. 2008. PMID: 18855426 Free PMC article.
-
Analyzing the catalytic role of Asp97 in the methionine aminopeptidase from Escherichia coli.FEBS J. 2008 Dec;275(24):6248-59. doi: 10.1111/j.1742-4658.2008.06749.x. Epub 2008 Nov 13. FEBS J. 2008. PMID: 19019076 Free PMC article.
-
Overexpression and divalent metal binding properties of the methionyl aminopeptidase from Pyrococcus furiosus.Biochemistry. 2002 Jun 11;41(23):7199-208. doi: 10.1021/bi020138p. Biochemistry. 2002. PMID: 12044150
-
Methionine as translation start signal: a review of the enzymes of the pathway in Escherichia coli.Biochimie. 1993;75(12):1061-75. doi: 10.1016/0300-9084(93)90005-d. Biochimie. 1993. PMID: 8199241 Review.
-
Structure and function of the methionine aminopeptidases.Biochim Biophys Acta. 2000 Mar 7;1477(1-2):157-67. doi: 10.1016/s0167-4838(99)00271-x. Biochim Biophys Acta. 2000. PMID: 10708856 Review.
References
-
- Meinnel T, Mechulam Y, Blanquet S. Methionine as translation start signal — a review of the enzymes of the pathway in Escherichia coli. Biochimie. 1993;75:1061–1075. - PubMed
-
- Arfin SM, Bradshaw RA. Cotranslational processing and protein turnover in eukaryotic cells. Biochemistry. 1988;27(21):7979–7984. - PubMed
-
- Bradshaw RA. Protein translation and turnover in eukaryotic cells. TIBS. 1989;14:276–279. - PubMed
-
- Bradshaw RA, Brickey WW, Walker KW. N-terminal processing: the methionine aminopeptidase and Nα-acetyl transferase families. TIBS. 1998;23:263–267. - PubMed
-
- Lowther WT, Matthews BW. Structure and function of the methionine aminopeptidases. Biochim. Biophys. Acta. 2000;1477:157–167. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

