Genome-wide screen in Francisella novicida for genes required for pulmonary and systemic infection in mice
- PMID: 18955478
- PMCID: PMC2612238
- DOI: 10.1128/IAI.00978-08
Genome-wide screen in Francisella novicida for genes required for pulmonary and systemic infection in mice
Abstract
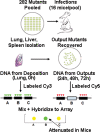
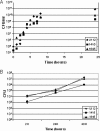
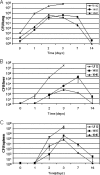
Francisella tularensis is a gram-negative, highly infectious, aerosolizable facultative intracellular pathogen that causes the potentially life-threatening disease tularemia. To date there is no approved vaccine available, and little is known about the molecular mechanisms important for infection, survival, and dissemination at different times of infection. We report the first whole-genome screen using an inhalation mouse model to monitor infection in the lung and dissemination to the liver and spleen. We queried a comprehensive library of 2,998 sequence-defined transposon insertion mutants in Francisella novicida strain U112 using a microarray-based negative-selection screen. We were able to track the behavior of 1,029 annotated genes, equivalent to a detection rate of 75% and corresponding to approximately 57% of the entire F. novicida genome. As expected, most transposon mutants retained the ability to colonize, but 125 candidate virulence genes (12%) could not be detected in at least one of the three organs. They fell into a variety of functional categories, with one-third having no annotated function and a statistically significant enrichment of genes involved in transcription. Based on the observation that behavior during complex pool infections correlated with the degree of attenuation during single-strain infection we identified nine genes expected to strongly contribute to infection. These included two genes, those for ATP synthase C (FTN_1645) and thioredoxin (FTN_1415), that when mutated allowed increased host survival and conferred protection in vaccination experiments.
Figures



Similar articles
-
TolC and EmrA1 contribute to Francisella novicida multidrug resistance and modulation of host cell death.J Bacteriol. 2024 Sep 19;206(9):e0024624. doi: 10.1128/jb.00246-24. Epub 2024 Aug 28. J Bacteriol. 2024. PMID: 39194223 Free PMC article.
-
Global Analysis of Genes Essential for Francisella tularensis Schu S4 Growth In Vitro and for Fitness during Competitive Infection of Fischer 344 Rats.J Bacteriol. 2019 Mar 13;201(7):e00630-18. doi: 10.1128/JB.00630-18. Print 2019 Apr 1. J Bacteriol. 2019. PMID: 30642993 Free PMC article.
-
Attenuated Francisella novicida transposon mutants protect mice against wild-type challenge.Infect Immun. 2006 Sep;74(9):5095-105. doi: 10.1128/IAI.00598-06. Infect Immun. 2006. PMID: 16926401 Free PMC article.
-
[Pathogenicity of Francisella].Zh Mikrobiol Epidemiol Immunobiol. 2005 Jan-Feb;(1):106-10. Zh Mikrobiol Epidemiol Immunobiol. 2005. PMID: 15773415 Review. Russian.
-
Importance of Metabolic Adaptations in Francisella Pathogenesis.Front Cell Infect Microbiol. 2017 Mar 28;7:96. doi: 10.3389/fcimb.2017.00096. eCollection 2017. Front Cell Infect Microbiol. 2017. PMID: 28401066 Free PMC article. Review.
Cited by
-
Prokaryotic Solute/Sodium Symporters: Versatile Functions and Mechanisms of a Transporter Family.Int J Mol Sci. 2021 Feb 13;22(4):1880. doi: 10.3390/ijms22041880. Int J Mol Sci. 2021. PMID: 33668649 Free PMC article. Review.
-
Working toward the future: insights into Francisella tularensis pathogenesis and vaccine development.Microbiol Mol Biol Rev. 2009 Dec;73(4):684-711. doi: 10.1128/MMBR.00028-09. Microbiol Mol Biol Rev. 2009. PMID: 19946137 Free PMC article. Review.
-
Francisella tularensis Schu S4 O-antigen and capsule biosynthesis gene mutants induce early cell death in human macrophages.Infect Immun. 2011 Feb;79(2):581-94. doi: 10.1128/IAI.00863-10. Epub 2010 Nov 15. Infect Immun. 2011. PMID: 21078861 Free PMC article.
-
Development, Strategies, and Challenges for Tularemia Vaccine.Curr Microbiol. 2024 Apr 2;81(5):126. doi: 10.1007/s00284-024-03658-0. Curr Microbiol. 2024. PMID: 38564047 Review.
-
An Expanded Transposon Mutant Library Reveals that Vibrio fischeri δ-Aminolevulinate Auxotrophs Can Colonize Euprymna scolopes.Appl Environ Microbiol. 2017 Feb 15;83(5):e02470-16. doi: 10.1128/AEM.02470-16. Print 2017 Mar 1. Appl Environ Microbiol. 2017. PMID: 28003196 Free PMC article.
References
-
- Boyer, P. D. 1997. The ATP synthase—a splendid molecular machine. Annu. Rev. Biochem. 66717-749. - PubMed
-
- Buchmeier, N. A., C. J. Lipps, M. Y. So, and F. Heffron. 1993. Recombination-deficient mutants of Salmonella typhimurium are avirulent and sensitive to the oxidative burst of macrophages. Mol. Microbiol. 7933-936. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

