Alternatively initiated gene 50/RTA transcripts expressed during murine and human gammaherpesvirus reactivation from latency
- PMID: 18971285
- PMCID: PMC2612301
- DOI: 10.1128/JVI.01444-08
Alternatively initiated gene 50/RTA transcripts expressed during murine and human gammaherpesvirus reactivation from latency
Abstract
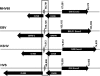
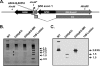
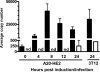
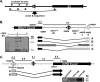
In the process of characterizing the requirements for expression of the essential immediate-early transcriptional activator (RTA) encoded by gene 50 of murine gammaherpesvirus 68 (MHV68), a recombinant virus was generated in which the known gene 50 promoter was deleted (G50pKO). Surprisingly, the G50pKO mutant retained the ability to replicate in permissive murine fibroblasts, albeit with slower kinetics than wild-type MHV68. 5'-rapid amplification of cDNA ends analyses of RNA prepared from G50pKO-infected fibroblasts revealed a novel upstream transcription initiation site, which was also utilized during wild-type MHV68 infection of permissive cells. Furthermore, the region upstream of the distal gene 50/RTA transcription initiation site exhibited promoter activity in both permissive NIH 3T12 fibroblasts as well as in the murine macrophage cell line RAW 264.7. In addition, in RAW 264.7 cells the activity of the distal gene 50/RTA promoter was strongly upregulated (>20-fold) by treatment of the cells with lipopolysaccharide. Reverse transcriptase PCR analyses of RNA prepared from Kaposi's sarcoma-associated herpesvirus- and Epstein-Barr virus-infected B-cell lines, following induction of virus reactivation, also revealed the presence of gene 50/RTA transcripts initiating upstream of the known transcription initiation site. The latter argues that alternative initiation of gene 50/RTA transcription is a strategy conserved among murine and human gammaherpesviruses. Infection of mice with the MHV68 G50pKO demonstrated the ability of this mutant virus to establish latency in the spleen and peritoneal exudate cells (PECs). However, the G50pKO mutant was unable to reactivate from latently infected splenocytes and also exhibited a significant reactivation defect from latently infected PECs, arguing in favor of a model where the proximal gene 50/RTA promoter plays a critical role in virus reactivation from latency, particularly from B cells. Finally, analyses of viral genome methylation in the regions upstream of the proximal and distal gene 50/RTA transcription initiation sites revealed that the distal promoter is partially methylated in vivo and heavily methylated in MHV68 latently infected B-cell lines, suggesting that DNA methylation may serve to silence the activity of this promoter during virus latency.
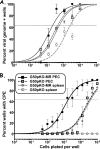
Figures






Similar articles
-
The de novo methyltransferases DNMT3a and DNMT3b target the murine gammaherpesvirus immediate-early gene 50 promoter during establishment of latency.J Virol. 2010 May;84(10):4946-59. doi: 10.1128/JVI.00060-10. Epub 2010 Mar 3. J Virol. 2010. PMID: 20200245 Free PMC article.
-
Identification of alternative transcripts encoding the essential murine gammaherpesvirus lytic transactivator RTA.J Virol. 2014 May;88(10):5474-90. doi: 10.1128/JVI.03110-13. Epub 2014 Feb 26. J Virol. 2014. PMID: 24574412 Free PMC article.
-
Identification of Novel Kaposi's Sarcoma-Associated Herpesvirus Orf50 Transcripts: Discovery of New RTA Isoforms with Variable Transactivation Potential.J Virol. 2016 Dec 16;91(1):e01434-16. doi: 10.1128/JVI.01434-16. Print 2017 Jan 1. J Virol. 2016. PMID: 27795414 Free PMC article.
-
Lytic cycle switches of oncogenic human gammaherpesviruses.Adv Cancer Res. 2007;97:81-109. doi: 10.1016/S0065-230X(06)97004-3. Adv Cancer Res. 2007. PMID: 17419942 Review.
-
The Rta/Orf50 transactivator proteins of the gamma-herpesviridae.Curr Top Microbiol Immunol. 2007;312:71-100. doi: 10.1007/978-3-540-34344-8_3. Curr Top Microbiol Immunol. 2007. PMID: 17089794 Review.
Cited by
-
Gammaherpesvirus gene expression and DNA synthesis are facilitated by viral protein kinase and histone variant H2AX.Virology. 2011 Nov 25;420(2):73-81. doi: 10.1016/j.virol.2011.08.019. Epub 2011 Sep 22. Virology. 2011. PMID: 21943826 Free PMC article.
-
The de novo methyltransferases DNMT3a and DNMT3b target the murine gammaherpesvirus immediate-early gene 50 promoter during establishment of latency.J Virol. 2010 May;84(10):4946-59. doi: 10.1128/JVI.00060-10. Epub 2010 Mar 3. J Virol. 2010. PMID: 20200245 Free PMC article.
-
Gammaherpesvirus-driven plasma cell differentiation regulates virus reactivation from latently infected B lymphocytes.PLoS Pathog. 2009 Nov;5(11):e1000677. doi: 10.1371/journal.ppat.1000677. Epub 2009 Nov 26. PLoS Pathog. 2009. PMID: 19956661 Free PMC article.
-
Redefining the genetics of murine gammaherpesvirus 68 via transcriptome-based annotation.Cell Host Microbe. 2010 Jun 25;7(6):516-26. doi: 10.1016/j.chom.2010.05.005. Cell Host Microbe. 2010. PMID: 20542255 Free PMC article.
-
RNA-guided gene editing of the murine gammaherpesvirus 68 genome reduces infectious virus production.PLoS One. 2021 Jun 4;16(6):e0252313. doi: 10.1371/journal.pone.0252313. eCollection 2021. PLoS One. 2021. PMID: 34086743 Free PMC article.
References
-
- Ben-Sasson, S. A., and G. Klein. 1981. Activation of the Epstein-Barr virus genome by 5-aza-cytidine in latently infected human lymphoid lines. Int. J. Cancer 28131-135. - PubMed
-
- Bhende, P. M., W. T. Seaman, H. J. Delecluse, and S. C. Kenney. 2004. The EBV lytic switch protein, Z, preferentially binds to and activates the methylated viral genome. Nat. Genet. 361099-1104. - PubMed
-
- Blaskovic, D., M. Stancekova, J. Svobodova, and J. Mistrikova. 1980. Isolation of five strains of herpesviruses from two species of free living small rodents. Acta Virol. 24468. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

