Novel long non-protein coding RNAs involved in Arabidopsis differentiation and stress responses
- PMID: 18997003
- PMCID: PMC2612962
- DOI: 10.1101/gr.080275.108
Novel long non-protein coding RNAs involved in Arabidopsis differentiation and stress responses
Abstract
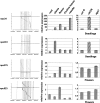
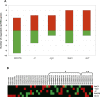
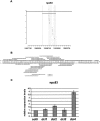
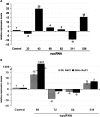
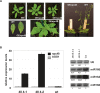
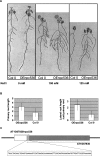
Long non-protein coding RNAs (npcRNA) represent an emerging class of riboregulators, which either act directly in this long form or are processed to shorter miRNA and siRNA. Genome-wide bioinformatic analysis of full-length cDNA databases identified 76 Arabidopsis npcRNAs. Fourteen npcRNAs were antisense to protein-coding mRNAs, suggesting cis-regulatory roles. Numerous 24-nt siRNA matched to five different npcRNAs, suggesting that these npcRNAs are precursors of this type of siRNA. Expression analyses of the 76 npcRNAs identified a novel npcRNA that accumulates in a dcl1 mutant but does not appear to produce trans-acting siRNA or miRNA. Additionally, another npcRNA was the precursor of miR869 and shown to be up-regulated in dcl4 but not in dcl1 mutants, indicative of a young miRNA gene. Abiotic stress altered the accumulation of 22 npcRNAs among the 76, a fraction significantly higher than that observed for the RNA binding protein-coding fraction of the transcriptome. Overexpression analyses in Arabidopsis identified two npcRNAs as regulators of root growth during salt stress and leaf morphology, respectively. Hence, together with small RNAs, long npcRNAs encompass a sensitive component of the transcriptome that have diverse roles during growth and differentiation.
Figures

References
-
- Adenot X., Elmayan T., Lauressergues D., Boutet S., Bouche N., Gasciolli V., Vaucheret H. DRB4-dependent TAS3 trans-acting siRNAs control leaf morphology through AGO7. Curr. Biol. 2006;16:927–932. - PubMed
-
- Argaman L., Hershberg R., Vogel J., Bejerano G., Wagner E.G., Margalit H., Altuvia S. Novel small RNA-encoding genes in the intergenic regions of Escherichia coli. Curr. Biol. 2001;11:941–950. - PubMed
-
- Bechtold N., Pelletier G. In planta Agrobacterium-mediated transformation of adult Arabidopsis thaliana plants by vacuum infiltration. Methods Mol. Biol. 1998;82:259–266. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
