Resolving genetic functions within microbial populations: in situ analyses using rRNA and mRNA stable isotope probing coupled with single-cell raman-fluorescence in situ hybridization
- PMID: 18997025
- PMCID: PMC2612214
- DOI: 10.1128/AEM.01861-08
Resolving genetic functions within microbial populations: in situ analyses using rRNA and mRNA stable isotope probing coupled with single-cell raman-fluorescence in situ hybridization
Abstract
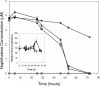
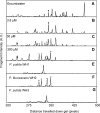
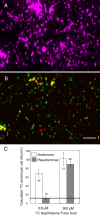
Prokaryotes represent one-half of the living biomass on Earth, with the vast majority remaining elusive to culture and study within the laboratory. As a result, we lack a basic understanding of the functions that many species perform in the natural world. To address this issue, we developed complementary population and single-cell stable isotope ((13)C)-linked analyses to determine microbial identity and function in situ. We demonstrated that the use of rRNA/mRNA stable isotope probing (SIP) recovered the key phylogenetic and functional RNAs. This was followed by single-cell physiological analyses of these populations to determine and quantify in situ functions within an aerobic naphthalene-degrading groundwater microbial community. Using these culture-independent approaches, we identified three prokaryote species capable of naphthalene biodegradation within the groundwater system: two taxa were isolated in the laboratory (Pseudomonas fluorescens and Pseudomonas putida), whereas the third eluded culture (an Acidovorax sp.). Using parallel population and single-cell stable isotope technologies, we were able to identify an unculturable Acidovorax sp. which played the key role in naphthalene biodegradation in situ, rather than the culturable naphthalene-biodegrading Pseudomonas sp. isolated from the same groundwater. The Pseudomonas isolates actively degraded naphthalene only at naphthalene concentrations higher than 30 muM. This study demonstrated that unculturable microorganisms could play important roles in biodegradation in the ecosystem. It also showed that the combined RNA SIP-Raman-fluorescence in situ hybridization approach may be a significant tool in resolving ecology, functionality, and niche specialization within the unculturable fraction of organisms residing in the natural environment.
Figures


Similar articles
-
A culture-independent approach to unravel uncultured bacteria and functional genes in a complex microbial community.PLoS One. 2012;7(10):e47530. doi: 10.1371/journal.pone.0047530. Epub 2012 Oct 17. PLoS One. 2012. PMID: 23082176 Free PMC article.
-
Raman-FISH: combining stable-isotope Raman spectroscopy and fluorescence in situ hybridization for the single cell analysis of identity and function.Environ Microbiol. 2007 Aug;9(8):1878-89. doi: 10.1111/j.1462-2920.2007.01352.x. Environ Microbiol. 2007. PMID: 17635536
-
Benzene Degradation by a Variovorax Species within a Coal Tar-Contaminated Groundwater Microbial Community.Appl Environ Microbiol. 2017 Feb 1;83(4):e02658-16. doi: 10.1128/AEM.02658-16. Print 2017 Feb 15. Appl Environ Microbiol. 2017. PMID: 27913419 Free PMC article.
-
Linking microbial community structure with function: fluorescence in situ hybridization-microautoradiography and isotope arrays.Curr Opin Biotechnol. 2006 Feb;17(1):83-91. doi: 10.1016/j.copbio.2005.12.006. Epub 2005 Dec 27. Curr Opin Biotechnol. 2006. PMID: 16377170 Review.
-
Unraveling uncultivable pesticide degraders via stable isotope probing (SIP).Crit Rev Biotechnol. 2018 Nov;38(7):1025-1048. doi: 10.1080/07388551.2018.1427697. Epub 2018 Feb 1. Crit Rev Biotechnol. 2018. PMID: 29385846 Review.
Cited by
-
Evident bacterial community changes but only slight degradation when polluted with pyrene in a red soil.Front Microbiol. 2015 Jan 30;6:22. doi: 10.3389/fmicb.2015.00022. eCollection 2015. Front Microbiol. 2015. PMID: 25688237 Free PMC article.
-
Revealing CO2-Fixing SAR11 Bacteria in the Ocean by Raman-Based Single-Cell Metabolic Profiling and Genomics.Biodes Res. 2022 Oct 13;2022:9782712. doi: 10.34133/2022/9782712. eCollection 2022. Biodes Res. 2022. PMID: 37850122 Free PMC article.
-
Capturing the genetic makeup of the active microbiome in situ.ISME J. 2017 Sep;11(9):1949-1963. doi: 10.1038/ismej.2017.59. Epub 2017 Jun 2. ISME J. 2017. PMID: 28574490 Free PMC article. Review.
-
A culture-independent approach to unravel uncultured bacteria and functional genes in a complex microbial community.PLoS One. 2012;7(10):e47530. doi: 10.1371/journal.pone.0047530. Epub 2012 Oct 17. PLoS One. 2012. PMID: 23082176 Free PMC article.
-
Mid-Infrared Photothermal-Fluorescence In Situ Hybridization for Functional Analysis and Genetic Identification of Single Cells.Anal Chem. 2023 Jan 31;95(4):2398-2405. doi: 10.1021/acs.analchem.2c04474. Epub 2023 Jan 18. Anal Chem. 2023. PMID: 36652555 Free PMC article.
References
-
- Bosch, R., E. Garcia-Valdes, and E. R. B. Moore. 2000. Complete nucleotide sequence and evolutionary significance of a chromosomally encoded naphthalene-degradation lower pathway from Pseudomonas stutzeri AN10. Gene 245:65-74. - PubMed
-
- Bosch, R., E. Garcia-Valdes, and E. R. B. Moore. 1999. Genetic characterization and evolutionary implications of a chromosomally encoded naphthalene-degradation upper pathway from Pseudomonas stutzeri AN10. Gene 236:149-157. - PubMed
-
- Cadisch, G., M. Espana, R. Causey, M. Richter, E. Shaw, J. A. W. Morgan, C. Rahn, and G. D. Bending. 2005. Technical considerations for the use of N-15-DNA stable-isotope probing for functional microbial activity in soils. Rapid Commun. Mass Spectrom. 19:1424-1428. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

