Notch signaling mediates G1/S cell-cycle progression in T cells via cyclin D3 and its dependent kinases
- PMID: 19001083
- PMCID: PMC2647664
- DOI: 10.1182/blood-2008-03-147967
Notch signaling mediates G1/S cell-cycle progression in T cells via cyclin D3 and its dependent kinases
Abstract
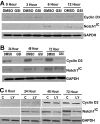
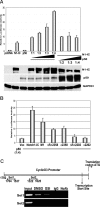
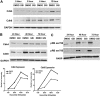
Notch signaling plays a role in normal lymphocyte development and function. Activating Notch1-mutations, leading to aberrant downstream signaling, have been identified in human T-cell acute lymphoblastic leukemia (T-ALL). While this highlights the contribution of Notch signaling to T-ALL pathogenesis, the mechanisms by which Notch regulates proliferation and survival in normal and leukemic T cells are not fully understood. Our findings identify a role for Notch signaling in G(1)-S progression of cell cycle in T cells. Here we show that expression of the G(1) proteins, cyclin D3, CDK4, and CDK6, is Notch-dependent both in vitro and in vivo, and we outline a possible mechanism for the regulated expression of cyclin D3 in activated T cells via CSL (CBF-1, mammals; suppressor of hairless, Drosophila melanogaster; Lag-1, Caenorhabditis elegans), as well as a noncanonical Notch signaling pathway. While cyclin D3 expression contributes to cell-cycle progression in Notch-dependent human T-ALL cell lines, ectopic expression of CDK4 or CDK6 together with cyclin D3 shows partial rescue from gamma-secretase inhibitor (GSI)-induced G(1) arrest in these cell lines. Importantly, cyclin D3 and CDK4 are highly overexpressed in Notch-dependent T-cell lymphomas, justifying the combined use of cell-cycle inhibitors and GSI in treating human T-cell malignancies.
Figures



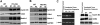
Comment in
-
Another Notch on the belt.Blood. 2009 Feb 19;113(8):1615-6. doi: 10.1182/blood-2008-12-192815. Blood. 2009. PMID: 19228927 No abstract available.
Similar articles
-
Inhibition of NOTCH signaling by gamma secretase inhibitor engages the RB pathway and elicits cell cycle exit in T-cell acute lymphoblastic leukemia cells.Cancer Res. 2009 Apr 1;69(7):3060-8. doi: 10.1158/0008-5472.CAN-08-4295. Epub 2009 Mar 24. Cancer Res. 2009. PMID: 19318552
-
KSHV viral cyclin interferes with T-cell development and induces lymphoma through Cdk6 and Notch activation in vivo.Cell Cycle. 2014;13(23):3670-84. doi: 10.4161/15384101.2014.964118. Cell Cycle. 2014. PMID: 25483078 Free PMC article.
-
NOTCH-1 and NOTCH-4 are novel gene targets of PEA3 in breast cancer: novel therapeutic implications.Breast Cancer Res. 2011 Jun 14;13(3):R63. doi: 10.1186/bcr2900. Breast Cancer Res. 2011. PMID: 21679465 Free PMC article.
-
New insights into Notch1 regulation of the PI3K-AKT-mTOR1 signaling axis: targeted therapy of γ-secretase inhibitor resistant T-cell acute lymphoblastic leukemia.Cell Signal. 2014 Jan;26(1):149-61. doi: 10.1016/j.cellsig.2013.09.021. Epub 2013 Oct 16. Cell Signal. 2014. PMID: 24140475 Review.
-
Cyclins and CDKS in development and cancer: lessons from genetically modified mice.Front Biosci. 2006 Jan 1;11:1164-88. doi: 10.2741/1871. Front Biosci. 2006. PMID: 16146805 Review.
Cited by
-
The NOTCH signaling pathway: role in the pathogenesis of T-cell acute lymphoblastic leukemia and implication for therapy.Ther Adv Hematol. 2013 Jun;4(3):199-210. doi: 10.1177/2040620712471368. Ther Adv Hematol. 2013. PMID: 23730497 Free PMC article.
-
Small Molecules in the Treatment of Squamous Cell Carcinomas: Focus on Indirubins.Cancers (Basel). 2021 Apr 7;13(8):1770. doi: 10.3390/cancers13081770. Cancers (Basel). 2021. PMID: 33917267 Free PMC article. Review.
-
miR-449a Repression Leads to Enhanced NOTCH Signaling in TMPRSS2:ERG Fusion Positive Prostate Cancer Cells.Cancers (Basel). 2021 Feb 25;13(5):964. doi: 10.3390/cancers13050964. Cancers (Basel). 2021. PMID: 33669024 Free PMC article.
-
Pegylated arginase I: a potential therapeutic approach in T-ALL.Blood. 2010 Jun 24;115(25):5214-21. doi: 10.1182/blood-2009-12-258822. Epub 2010 Apr 20. Blood. 2010. PMID: 20407034 Free PMC article.
-
Oncogenic transcriptional program driven by TAL1 in T-cell acute lymphoblastic leukemia.Int J Hematol. 2019 Jan;109(1):5-17. doi: 10.1007/s12185-018-2518-z. Epub 2018 Aug 25. Int J Hematol. 2019. PMID: 30145780 Review.
References
-
- Artavanis-Tsakonas S, Rand MD, Lake RJ. Notch signaling: cell fate control and signal integration in development. Science. 1999;284:770–776. - PubMed
-
- Radtke F, Wilson A, Mancini SJC, MacDonald HR. Notch regulation of lymphocyte development and function. Nat Immunol. 2004;5:247–253. - PubMed
-
- Maillard I, Fang T, Pear WS. Regulation of lymphoid development, differentiation, and function by the Notch pathway. Annu Rev Immunol. 2005;23:945–974. - PubMed
-
- Aster JC. Deregulated NOTCH signaling in acute T-cell lymphoblastic leukemia/lymphoma: new insights, questions, and opportunities. Int J Hematol. 2005;82:295–301. - PubMed
-
- Grabher C, von Boehmer H, Look AT. Notch 1 activation in the molecular pathogenesis of T-cell acute lymphoblastic leukaemia. Nat Rev Cancer. 2006;6:347–359. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

