High nucleotide divergence in developmental regulatory genes contrasts with the structural elements of olfactory pathways in caenorhabditis
- PMID: 19001295
- PMCID: PMC2666507
- DOI: 10.1534/genetics.107.082651
High nucleotide divergence in developmental regulatory genes contrasts with the structural elements of olfactory pathways in caenorhabditis
Abstract
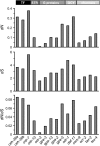
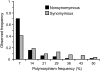
Almost all organismal function is controlled by pathways composed of interacting genetic components. The relationship between pathway structure and the evolution of individual pathway components is not completely understood. For the nematode Caenorhabditis elegans, chemosensory pathways regulate critical aspects of an individual's life history and development. To help understand how olfaction evolves in Caenorhabditis and to examine patterns of gene evolution within transduction pathways in general, we analyzed nucleotide variation within and between species across two well-characterized olfactory pathways, including regulatory genes controlling the fate of the cells in which the pathways are expressed. In agreement with previous studies, we found much higher levels of polymorphism within C. remanei than within the related species C. elegans and C. briggsae. There are significant differences in the rates of nucleotide evolution for genes across the two pathways but no particular association between evolutionary rate and gene position, suggesting that the evolution of functional pathways must be considered within the context of broader gene network structure. However, developmental regulatory genes show both higher levels of divergence and polymorphism than the structural genes of the pathway. These results show that, contrary to the emerging paradigm in the evolution of development, important structural changes can accumulate in transcription factors.
Figures

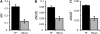
References
-
- Ache, B. W., and J. M. Young, 2005. Olfaction: diverse species, conserved principles. Neuron 48 417–430. - PubMed
-
- Altschul, S. F., W. Gish, W. Miller, E. W. Myers and D. J. Lipman, 1990. Basic local alignment search tool. J. Mol. Biol. 215 403–410. - PubMed
-
- Baird, S. E., 1999. Natural and experimental associations of Caenorhabditis remanei with Trachelipus rathkii and other terrestrial isopods. Nematology 1 471–475.
-
- Bargmann, C. I., E. Hartwieg and H. R. Horvitz, 1993. Odorant-selective genes and neurons mediate olfaction in C. elegans. Cell 74 515–527. - PubMed
Publication types
MeSH terms
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

