Distribution of label-retaining cells in the limbal epithelium of a mouse eye
- PMID: 19001638
- PMCID: PMC2628319
- DOI: 10.1369/jhc.2008.952390
Distribution of label-retaining cells in the limbal epithelium of a mouse eye
Abstract
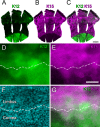
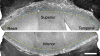
Corneal epithelial stem cells are believed to be localized in the limbus, an annular zone between the cornea and the conjunctiva, but it has not been possible to identify individual stem cells in situ because of the lack of specific molecular markers. Description of stem cell distribution has also been ambiguous because limbal boundaries are ill defined. In this study, we investigated whether distribution of slow cycling, label-retaining cells (LRCs) could be determined precisely against a definable anatomical structure of an eye. We found that a boundary between the cornea and the limbus could be determined reliably by distinct epithelial nuclear staining patterns. Using this boundary line as a fiduciary marker, we determined that LRCs were located exclusively in the basal epithelium at the limbal side of the cornea-limbus boundary line along the entire circumference, within an annular zone of 100-200 mum wide. LRC density was highest in the superior temporal quadrant and lowest in the inferior nasal quadrant. These results show that LRCs are present asymmetrically in a narrow zone within the limbus that can be defined precisely in reference to a newly defined anatomical boundary line between the cornea and the limbus.
Figures



Similar articles
-
Integrins in slow-cycling corneal epithelial cells at the limbus in the mouse.Stem Cells. 2006 Apr;24(4):1075-86. doi: 10.1634/stemcells.2005-0382. Epub 2005 Nov 10. Stem Cells. 2006. PMID: 16282441
-
Rare corneal clones in mice suggest an age-related decrease of stem cell activity and support the limbal epithelial stem cell hypothesis.Stem Cell Res. 2012 Jan;8(1):109-19. doi: 10.1016/j.scr.2011.08.007. Epub 2011 Sep 5. Stem Cell Res. 2012. PMID: 22099025
-
Limbal epithelial crypt: a model for corneal epithelial maintenance and novel limbal regional variations.Arch Ophthalmol. 2008 May;126(5):665-9. doi: 10.1001/archopht.126.5.665. Arch Ophthalmol. 2008. PMID: 18474777
-
Corneal epithelial stem cells at the limbus: looking at some old problems from a new angle.Exp Eye Res. 2004 Mar;78(3):433-46. doi: 10.1016/j.exer.2003.09.008. Exp Eye Res. 2004. PMID: 15106923 Review.
-
Limbal stem cells: the search for a marker.Clin Exp Ophthalmol. 2006 Jan-Feb;34(1):64-73. doi: 10.1111/j.1442-9071.2006.01147.x. Clin Exp Ophthalmol. 2006. PMID: 16451261 Review.
Cited by
-
Cellular and molecular profiles of larval and adult Xenopus corneal epithelia resolved at the single-cell level.Dev Biol. 2022 Nov;491:13-30. doi: 10.1016/j.ydbio.2022.08.007. Epub 2022 Aug 29. Dev Biol. 2022. PMID: 36049533 Free PMC article.
-
The lens regenerative competency of limbal vs. central regions of mature Xenopus cornea epithelium.Exp Eye Res. 2016 Nov;152:94-99. doi: 10.1016/j.exer.2016.08.013. Epub 2016 Aug 26. Exp Eye Res. 2016. PMID: 27569373 Free PMC article. Review.
-
Mosaic analysis of stem cell function and wound healing in the mouse corneal epithelium.BMC Dev Biol. 2009 Jan 7;9:4. doi: 10.1186/1471-213X-9-4. BMC Dev Biol. 2009. PMID: 19128502 Free PMC article.
-
Intracellular Position of Centrioles and the Direction of Homeostatic Epithelial Cell Movements in the Mouse Cornea.J Histochem Cytochem. 2017 Feb;65(2):83-91. doi: 10.1369/0022155416674718. Epub 2016 Nov 12. J Histochem Cytochem. 2017. PMID: 28117631 Free PMC article.
-
Computer simulation of neutral drift among limbal epithelial stem cells of mosaic mice.Stem Cell Res. 2018 Jul;30:1-11. doi: 10.1016/j.scr.2018.05.003. Epub 2018 May 13. Stem Cell Res. 2018. PMID: 29777801 Free PMC article.
References
-
- Beebe DC, Masters BR (1996) Cell lineage and the differentiation of corneal epithelial cells. Invest Ophthalmol Vis Sci 37:1815–1825 - PubMed
-
- Bickenbach JR (1981) Identification and behavior of label-retaining cells in oral mucosa and skin. J Dent Res 60:1611–1620 - PubMed
-
- Braun KM, Watt FM (2004) Epidermal label-retaining cells: background and recent applications. J Investig Dermatol Symp Proc 9:196–201 - PubMed
-
- Buck RC (1985) Measurement of centripetal migration of normal corneal epithelial cells in the mouse. Invest Ophthalmol Vis Sci 26:1296–1299 - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

