Concepts of epigenetics in prostate cancer development
- PMID: 19002169
- PMCID: PMC2634711
- DOI: 10.1038/sj.bjc.6604771
Concepts of epigenetics in prostate cancer development
Abstract
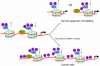
Substantial evidence now supports the view that epigenetic changes have a role in the development of human prostate cancer. Analyses of the patterns of epigenetic alteration are providing important insights into the origin of this disease and have identified specific alterations that may serve as useful diagnostic and prognostic biomarkers. Examination of cancer methylation patterns supports a stem cell origin of prostate cancer. It is well established that methylation of GSTpi is a marker of prostate cancer, and global patterns of histone marking appear to be linked to cancer prognosis with levels of acetylated histones H3K9, H3K18, and H4K12, and of dimethylated H4R3 and H3K4, dividing low-grade prostate cancer (Gleason 6 or less) into two prognostically separate groups. Elevated levels of several components of the polycomb group protein complex, EZH2, BMI1, and RING1, can also act as biomarkers of poor clinical outcome. Many components of the epigenetic machinery, including histone deacetylase (whose expression level is linked to the TMPRSS2:ERG translocation) and the histone methylase EZH2, are potential therapeutic targets. The recent discovery of the role of small RNAs in governing the epigenetic status of individual genes offers exciting new possibilities in therapeutics and chemoprevention.
Figures
References
-
- Beke L, Nuytten M, Van Eynde A, Beullens M, Bollen M (2007) The gene encoding the prostatic tumor suppressor PSP94 is a target for repression by the Polycomb group protein EZH2. Oncogene 26: 4590–4595 - PubMed
-
- Chen H, Tu SW, Hsieh JT (2005) Down-regulation of human DAB2IP gene expression mediated by polycomb Ezh2 complex and histone deacetylase in prostate cancer. J Biol Chem 280: 22437–22444 - PubMed
-
- Fraga MF, Ballestar E, Paz MF, Ropero S, Setien F, Ballestar ML, Heine-Suner D, Cigudosa JC, Urioste M, Benitez J, Boix-Chornet M, Sanchez-Aguilera A, Ling C, Carlsson E, Poulsen P, Vaag A, Stephan Z, Spector TD, Wu YZ, Plass C, Esteller M (2005a) Epigenetic differences arise during the lifetime of monozygotic twins. Proc Natl Acad Sci USA 102: 10604–10609 - PMC - PubMed
-
- Fraga MF, Ballestar E, Villar-Garea A, Boix-Chornet M, Espada J, Schotta G, Bonaldi T, Haydon C, Ropero S, Petrie K, Iyer NG, Perez-Rosado A, Calvo E, Lopez JA, Cano A, Calasanz MJ, Colomer D, Piris MA, Ahn N, Imhof A, Caldas C, Jenuwein T, Esteller M (2005b) Loss of acetylation at Lys16 and trimethylation at Lys20 of histone H4 is a common hallmark of human cancer. Nat Genet 37: 391–400 - PubMed
-
- Gallinari P, Di Marco S, Jones P, Pallaoro M, Steinkuhler C (2007) HDACs, histone deacetylation and gene transcription: from molecular biology to cancer therapeutics. Cell Res 17: 195–211 - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

