Highly L and D enantioselective variants of horseradish peroxidase discovered by an ultrahigh-throughput selection method
- PMID: 19004779
- PMCID: PMC2584688
- DOI: 10.1073/pnas.0809851105
Highly L and D enantioselective variants of horseradish peroxidase discovered by an ultrahigh-throughput selection method
Abstract
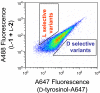
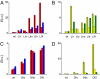
A highly efficient selection method for enhanced enzyme enantioselectivity based on yeast surface display and fluorescence-activated cell sorting (FACS) is developed and validated. Its application to horseradish peroxidase has resulted in enzyme variants up to 2 orders of magnitude selective toward either substrate enantiomer at will. These marked improvements in enantioselectivity are demonstrated for the surface-bound and soluble enzymes and rationalized by computational docking studies.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

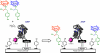
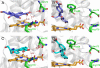
Similar articles
-
How a single-point mutation in horseradish peroxidase markedly enhances enantioselectivity.J Am Chem Soc. 2009 Aug 12;131(31):11155-60. doi: 10.1021/ja903482u. J Am Chem Soc. 2009. PMID: 19610634 Free PMC article.
-
Selection of horseradish peroxidase variants with enhanced enantioselectivity by yeast surface display.Chem Biol. 2007 Oct;14(10):1176-85. doi: 10.1016/j.chembiol.2007.09.008. Chem Biol. 2007. PMID: 17961829
-
Enantioselective oxidations of sulfides catalyzed by chloroperoxidase.Biochemistry. 1990 Nov 20;29(46):10465-8. doi: 10.1021/bi00498a006. Biochemistry. 1990. PMID: 2271658
-
Ultrahigh-throughput FACS-based screening for directed enzyme evolution.Chembiochem. 2009 Nov 23;10(17):2704-15. doi: 10.1002/cbic.200900384. Chembiochem. 2009. PMID: 19780076 Review.
-
Horseradish peroxidase: a modern view of a classic enzyme.Phytochemistry. 2004 Feb;65(3):249-59. doi: 10.1016/j.phytochem.2003.10.022. Phytochemistry. 2004. PMID: 14751298 Review.
Cited by
-
Cell wall trapping of autocrine peptides for human G-protein-coupled receptors on the yeast cell surface.PLoS One. 2012;7(5):e37136. doi: 10.1371/journal.pone.0037136. Epub 2012 May 18. PLoS One. 2012. PMID: 22623985 Free PMC article.
-
A High-Throughput Screening System Based on Droplet Microfluidics for Glucose Oxidase Gene Libraries.Molecules. 2020 May 22;25(10):2418. doi: 10.3390/molecules25102418. Molecules. 2020. PMID: 32455903 Free PMC article.
-
How a single-point mutation in horseradish peroxidase markedly enhances enantioselectivity.J Am Chem Soc. 2009 Aug 12;131(31):11155-60. doi: 10.1021/ja903482u. J Am Chem Soc. 2009. PMID: 19610634 Free PMC article.
-
Beyond antibody engineering: directed evolution of alternative binding scaffolds and enzymes using yeast surface display.Microb Cell Fact. 2018 Feb 26;17(1):32. doi: 10.1186/s12934-018-0881-3. Microb Cell Fact. 2018. PMID: 29482656 Free PMC article. Review.
-
High-throughput screening of microchip-synthesized genes in programmable double-emulsion droplets.Nanoscale. 2017 Mar 9;9(10):3485-3495. doi: 10.1039/c6nr08224f. Nanoscale. 2017. PMID: 28239692 Free PMC article.
References
-
- Rouhi M. Chiral chemistry: Traditional methods thrive despite numerous hurdles, including tough luck, slow commercialization of catalytic processes. Chem Eng News. 2004;82:47–62.
-
- Shin H-D, Guo X, Chen R. In: Bioprocessing for Value Added Products from Renewable Resources. Yang ST, editor. Amsterdam: Elsevier; 2006. pp. 351–371.
-
- Klibanov AM. Improving enzymes by using them in organic solvents. Nature. 2001;409:241–246. - PubMed
-
- Klibanov AM. Enzyme memory—What is remembered and why? Nature. 1995;374:596. - PubMed
-
- Phillips RS. Temperature modulation of the stereochemistry of enzymatic catalysis: Prospects for exploitation. Trends Biotechnol. 1996;14:13–16.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

