The EPHA2 gene is associated with cataracts linked to chromosome 1p
- PMID: 19005574
- PMCID: PMC2582197
The EPHA2 gene is associated with cataracts linked to chromosome 1p
Abstract
Purpose: Cataracts are a clinically and genetically heterogeneous disorder affecting the ocular lens, and the leading cause of treatable vision loss and blindness worldwide. Here we identify a novel gene linked with a rare autosomal dominant form of childhood cataracts segregating in a four generation pedigree, and further show that this gene is likely associated with much more common forms of age-related cataracts in a case-control cohort.
Methods: Genomic DNA was prepared from blood leukocytes, and genotyping was performed by means of single nucleotide polymorphism (SNP) markers, and short tandem repeat (STR) markers. Linkage analyses were performed with the GeneHunter and MLINK programs, and association analyses were performed with the Haploview and Exemplar programs. Mutation detection was achieved by PCR amplification of exons and di-deoxy cycle-sequencing.
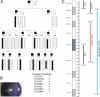
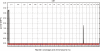
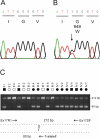
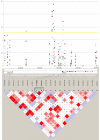
Results: Genome-wide linkage analysis with SNP markers, identified a likely disease-haplotype interval on chromosome 1p (rs707455-[approximately 10 Mb]-rs477558). Linkage to chromosome 1p was confirmed using STR markers D1S2672 (LOD score [Z]=3.56, recombination distance [theta]=0), and D1S2697 (Z=2.92, theta=0). Mutation profiling of positional-candidate genes detected a heterozygous transversion (c.2842G>T) in exon 17 of the gene coding for Eph-receptor type-A2 (EPHA2) that cosegregated with the disease. This missense change was predicted to result in the non-conservative substitution of a tryptophan residue for a phylogenetically conserved glycine residue at codon 948 (p.G948W), within a conserved cytoplasmic domain of the receptor. Candidate gene association analysis further identified SNPs in the EPHA2 region of chromosome 1p that were suggestively associated with age-related cataracts (p=0.007 for cortical cataracts, and p=0.01 for cortical and/or nuclear cataracts).
Conclusions: These data provide the first evidence that EPHA2, which functions in the Eph-ephrin bidirectional signaling pathway of mammalian cells, plays a vital role in maintaining lens transparency.
Figures
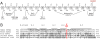
References
-
- World Health Organization - Vision 2020; The Right to Sight, Global initiative for the elimination of avoidable blindness - Action plan 2006–2011 (http://www.who.int/blindness/Vision2020%20-report.pdf)
-
- SanGiovanni JP. Infantile cataract in the collaborative perinatal project: prevalence and risk factors. Arch Ophthalmol. 2002;120:1559–65. - PubMed
-
- Zetterstrom C. Cataracts in children. J Cataract Refract Surg. 2005;31:824–40. - PubMed
-
- Shiels A. Genetic origins of cataract. Arch Ophthalmol. 2007;125:165–73. - PubMed
-
- Litt M. Autosomal dominant congenital cataract associated with a missense mutation in the human alpha crystallin gene CRYAA. Hum Mol Genet. 1998;7:471–4. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous
