Dissection of a QTL hotspot on mouse distal chromosome 1 that modulates neurobehavioral phenotypes and gene expression
- PMID: 19008955
- PMCID: PMC2577893
- DOI: 10.1371/journal.pgen.1000260
Dissection of a QTL hotspot on mouse distal chromosome 1 that modulates neurobehavioral phenotypes and gene expression
Abstract
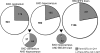
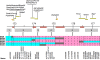
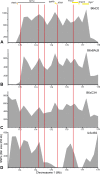
A remarkably diverse set of traits maps to a region on mouse distal chromosome 1 (Chr 1) that corresponds to human Chr 1q21-q23. This region is highly enriched in quantitative trait loci (QTLs) that control neural and behavioral phenotypes, including motor behavior, escape latency, emotionality, seizure susceptibility (Szs1), and responses to ethanol, caffeine, pentobarbital, and haloperidol. This region also controls the expression of a remarkably large number of genes, including genes that are associated with some of the classical traits that map to distal Chr 1 (e.g., seizure susceptibility). Here, we ask whether this QTL-rich region on Chr 1 (Qrr1) consists of a single master locus or a mixture of linked, but functionally unrelated, QTLs. To answer this question and to evaluate candidate genes, we generated and analyzed several gene expression, haplotype, and sequence datasets. We exploited six complementary mouse crosses, and combed through 18 expression datasets to determine class membership of genes modulated by Qrr1. Qrr1 can be broadly divided into a proximal part (Qrr1p) and a distal part (Qrr1d), each associated with the expression of distinct subsets of genes. Qrr1d controls RNA metabolism and protein synthesis, including the expression of approximately 20 aminoacyl-tRNA synthetases. Qrr1d contains a tRNA cluster, and this is a functionally pertinent candidate for the tRNA synthetases. Rgs7 and Fmn2 are other strong candidates in Qrr1d. FMN2 protein has pronounced expression in neurons, including in the dendrites, and deletion of Fmn2 had a strong effect on the expression of few genes modulated by Qrr1d. Our analysis revealed a highly complex gene expression regulatory interval in Qrr1, composed of multiple loci modulating the expression of functionally cognate sets of genes.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures





References
-
- DeFries JC, Gervais MC, Thomas EA. Response to 30 generations of selection for open-field activity in laboratory mice. Behav Genet. 1978;129:1432–1435. - PubMed
-
- Caldarone B, Saavedra C, Tartaglia K, Wehner JM, Dudek BC, et al. Quantitative trait loci analysis affecting contextual conditioning in mice. Nat Genet. 1997;17:335–337. - PubMed
-
- Gershenfeld HK, Neumann PE, Mathis C, Crawley JN, Li X, et al. Mapping quantitative trait loci for open-field behavior in mice. Behav Genet. 1997;27:201–210. - PubMed
-
- Flint J, Corley R, DeFries JC, Fulker DW, Gray JA, et al. A simple genetic basis for a complex psychological trait in laboratory mice. Science. 1995;269:1432–1435. - PubMed
-
- Turri MG, Talbot CJ, Radcliffe RA, Wehner JM, Flint J. High-resolution mapping of quantitative trait loci for emotionality in selected strains of mice. Mamm Genome. 1999;10:1098–1101. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

