Proteomic profiling of amniotic fluid in preterm labor using two-dimensional liquid separation and mass spectrometry
- PMID: 19012186
- PMCID: PMC3163445
- DOI: 10.1080/14767050802053289
Proteomic profiling of amniotic fluid in preterm labor using two-dimensional liquid separation and mass spectrometry
Abstract
Objective: Simultaneous analysis of the protein composition of biological fluids is now possible. Such an approach can be used to identify biological markers of disease and to understand the pathophysiology of disorders that have eluded classification, diagnosis, and treatment. The purpose of this study was to analyze the differences in protein composition of the amniotic fluid of patients in preterm labor.
Study design: Amniotic fluid was obtained by amniocentesis from three groups of women with preterm labor and intact membranes: (1) women without intra-amniotic infection/inflammation (IAI) who delivered at term, (2) women without IAI who delivered a preterm neonate, and (3) women with IAI. Intra-amniotic infection was defined as a positive amniotic fluid culture for microorganisms. Intra-amniotic inflammation was defined as an elevated amniotic fluid interleukin (IL)-6 (> or =2.3 ng/mL). Two-dimensional (2D) chromatography was used for analysis. The first dimension separated proteins by isoelectric point, while the second, by the degree of hydrophobicity. 2D protein maps were generated using different experimental conditions (reducing agents as well as protein concentration). The maps were used to discern subsets of isoelectric point/hydrophobicity containing differentially expressed proteins. Protein identification of differentially expressed fractions was conducted with mass spectrometry. Enzyme-linked immunosorbent assays (ELISA) as well as surface-enhanced laser desorption/ionization time-of-flight mass spectrometry (SELDI-TOF MS)-based on-chip antibody capture immunoassays were also used for confirmation of a specific protein that was differentially expressed.
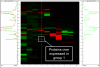
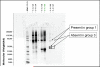
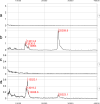
Results: (1) Amniotic fluid protein composition can be analyzed using a combination of 2D liquid chromatography and mass spectrometry for the identification of proteins differentially expressed in patients in preterm labor. (2) While total insulin-like growth factor-binding protein-1 (IGFBP-1) concentration did not change, IGFBP-1 fragments at about 13.5 kDa were present in patients with IAI. (3) Proteins that were over-expressed in group 1 included von Ebner gland protein precursor, IL-7 precursor, apolipoprotein A1, tropomyosin sk1 (TPMsk1) fragment, ribosomal protein S6 kinase alpha-3, and alpha-1-microglobulin/bikunin precursor (AMBP). (4) Proteins that were over-expressed in group 3 included fibrinopeptide B, transferrin, major histocompatibility complex (MHC) class 1 chain-related A antigen fragment, transcription elongation factor A, sex-determining region Y (SRY) box 5 protein, Down syndrome critical region 2 protein (DSCR2), and human peptide 8 (HP8). (5) One protein, retinol-binding protein, was over-expressed in women who delivered preterm, regardless of the presence of IAI.
Conclusions: A combination of techniques involving 2D chromatography, mass spectrometry, and immunoassays allows identification of proteins that are differentially regulated in the amniotic fluid of patients with preterm labor. Specifically, the amount of the IGFBP-1 fragments at approximately 13.5 kDa was found to be increased in patients with IAI, while the amount of the intact form of IGFBP-1 was decreased.
Figures





Similar articles
-
Isobaric labeling and tandem mass spectrometry: a novel approach for profiling and quantifying proteins differentially expressed in amniotic fluid in preterm labor with and without intra-amniotic infection/inflammation.J Matern Fetal Neonatal Med. 2010 Apr;23(4):261-80. doi: 10.3109/14767050903067386. J Matern Fetal Neonatal Med. 2010. PMID: 19670042 Free PMC article.
-
Diagnosis of intra-amniotic infection by proteomic profiling and identification of novel biomarkers.JAMA. 2004 Jul 28;292(4):462-9. doi: 10.1001/jama.292.4.462. JAMA. 2004. PMID: 15280344
-
Proteomic analysis of amniotic fluid to identify women with preterm labor and intra-amniotic inflammation/infection: the use of a novel computational method to analyze mass spectrometric profiling.J Matern Fetal Neonatal Med. 2008 Jun;21(6):367-88. doi: 10.1080/14767050802045848. J Matern Fetal Neonatal Med. 2008. PMID: 18570116 Free PMC article.
-
The use of high-dimensional biology (genomics, transcriptomics, proteomics, and metabolomics) to understand the preterm parturition syndrome.BJOG. 2006 Dec;113 Suppl 3(Suppl 3):118-35. doi: 10.1111/j.1471-0528.2006.01150.x. BJOG. 2006. PMID: 17206980 Free PMC article. Review.
-
Using proteomics in perinatal and neonatal sepsis: hopes and challenges for the future.Curr Opin Infect Dis. 2009 Jun;22(3):235-43. doi: 10.1097/QCO.0b013e32832a5963. Curr Opin Infect Dis. 2009. PMID: 19395960 Free PMC article. Review.
Cited by
-
Retinol binding protein 4: an adipokine associated with intra-amniotic infection/inflammation.J Matern Fetal Neonatal Med. 2010 Feb;23(2):111-9. doi: 10.3109/14767050902994739. J Matern Fetal Neonatal Med. 2010. PMID: 19900011 Free PMC article.
-
Human β-defensin-3 participates in intra-amniotic host defense in women with labor at term, spontaneous preterm labor and intact membranes, and preterm prelabor rupture of membranes.J Matern Fetal Neonatal Med. 2020 Dec;33(24):4117-4132. doi: 10.1080/14767058.2019.1597047. Epub 2019 Apr 18. J Matern Fetal Neonatal Med. 2020. PMID: 30999788 Free PMC article.
-
Metabolomics in premature labor: a novel approach to identify patients at risk for preterm delivery.J Matern Fetal Neonatal Med. 2010 Dec;23(12):1344-59. doi: 10.3109/14767058.2010.482618. Epub 2010 May 26. J Matern Fetal Neonatal Med. 2010. PMID: 20504069 Free PMC article.
-
The frequency and clinical significance of intra-amniotic inflammation in twin pregnancies with preterm labor and intact membranes.J Matern Fetal Neonatal Med. 2019 Feb;32(4):527-541. doi: 10.1080/14767058.2017.1384460. Epub 2017 Oct 11. J Matern Fetal Neonatal Med. 2019. PMID: 29020827 Free PMC article.
-
Amniotic fluid: the use of high-dimensional biology to understand fetal well-being.Reprod Sci. 2014 Jan;21(1):6-19. doi: 10.1177/1933719113485292. Epub 2013 Apr 18. Reprod Sci. 2014. PMID: 23599373 Free PMC article. Review.
References
-
- Evans GA. Designer science and the “omic” revolution. Nat.Biotechnol. 2000;18:127. - PubMed
-
- Gracey AY, Cossins AR. Application of microarray technology in environmental and comparative physiology. Annu.Rev.Physiol. 2003;65:231–259. - PubMed
-
- Vuadens F, Benay C, Crettaz D, Gallot D, Sapin V, Schneider P, Bienvenut WV, Lemery D, Quadroni M, Dastugue B, et al. Identification of biologic markers of the premature rupture of fetal membranes: proteomic approach. Proteomics. 2003;3:1521–1525. - PubMed
-
- Mehta T, Tanik M, Allison DB. Towards sound epistemological foundations of statistical methods for high-dimensional biology. Nat.Genet. 2004;36:943–947. - PubMed
-
- Strange K. The end of “naive reductionism”: rise of systems biology or renaissance of physiology? Am.J.Physiol Cell Physiol. 2005;288:C968–C974. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous
