A salmonid EST genomic study: genes, duplications, phylogeny and microarrays
- PMID: 19014685
- PMCID: PMC2628678
- DOI: 10.1186/1471-2164-9-545
A salmonid EST genomic study: genes, duplications, phylogeny and microarrays
Abstract
Background: Salmonids are of interest because of their relatively recent genome duplication, and their extensive use in wild fisheries and aquaculture. A comprehensive gene list and a comparison of genes in some of the different species provide valuable genomic information for one of the most widely studied groups of fish.
Results: 298,304 expressed sequence tags (ESTs) from Atlantic salmon (69% of the total), 11,664 chinook, 10,813 sockeye, 10,051 brook trout, 10,975 grayling, 8,630 lake whitefish, and 3,624 northern pike ESTs were obtained in this study and have been deposited into the public databases. Contigs were built and putative full-length Atlantic salmon clones have been identified. A database containing ESTs, assemblies, consensus sequences, open reading frames, gene predictions and putative annotation is available. The overall similarity between Atlantic salmon ESTs and those of rainbow trout, chinook, sockeye, brook trout, grayling, lake whitefish, northern pike and rainbow smelt is 93.4, 94.2, 94.6, 94.4, 92.5, 91.7, 89.6, and 86.2% respectively. An analysis of 78 transcript sets show Salmo as a sister group to Oncorhynchus and Salvelinus within Salmoninae, and Thymallinae as a sister group to Salmoninae and Coregoninae within Salmonidae. Extensive gene duplication is consistent with a genome duplication in the common ancestor of salmonids. Using all of the available EST data, a new expanded salmonid cDNA microarray of 32,000 features was created. Cross-species hybridizations to this cDNA microarray indicate that this resource will be useful for studies of all 68 salmonid species.
Conclusion: An extensive collection and analysis of salmonid RNA putative transcripts indicate that Pacific salmon, Atlantic salmon and charr are 94-96% similar while the more distant whitefish, grayling, pike and smelt are 93, 92, 89 and 86% similar to salmon. The salmonid transcriptome reveals a complex history of gene duplication that is consistent with an ancestral salmonid genome duplication hypothesis. Genome resources, including a new 32 K microarray, provide valuable new tools to study salmonids.
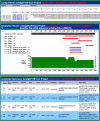
Figures


Similar articles
-
Salmonidae Genome: Features, Evolutionary and Phylogenetic Characteristics.Genes (Basel). 2022 Nov 27;13(12):2221. doi: 10.3390/genes13122221. Genes (Basel). 2022. PMID: 36553488 Free PMC article. Review.
-
Development and application of a salmonid EST database and cDNA microarray: data mining and interspecific hybridization characteristics.Genome Res. 2004 Mar;14(3):478-90. doi: 10.1101/gr.1687304. Epub 2004 Feb 12. Genome Res. 2004. PMID: 14962987 Free PMC article.
-
Characterization of the Atlantic salmon (Salmo salar) brain-type fatty acid binding protein (fabp7) genes reveals the fates of teleost fabp7 genes following whole genome duplications.Gene. 2012 Aug 10;504(2):253-61. doi: 10.1016/j.gene.2012.04.089. Epub 2012 May 8. Gene. 2012. PMID: 22575613
-
Genome evolution in the fish family salmonidae: generation of a brook charr genetic map and comparisons among charrs (Arctic charr and brook charr) with rainbow trout.BMC Genet. 2011 Jul 28;12:68. doi: 10.1186/1471-2156-12-68. BMC Genet. 2011. PMID: 21798024 Free PMC article.
-
The salmonid class I MHC: limited diversity in a primitive teleost.Immunol Rev. 1998 Dec;166:279-93. doi: 10.1111/j.1600-065x.1998.tb01269.x. Immunol Rev. 1998. PMID: 9914919 Review.
Cited by
-
Genomics in eels--towards aquaculture and biology.Mar Biotechnol (NY). 2012 Oct;14(5):583-90. doi: 10.1007/s10126-012-9444-5. Epub 2012 Apr 17. Mar Biotechnol (NY). 2012. PMID: 22527267 Free PMC article.
-
Salmonidae Genome: Features, Evolutionary and Phylogenetic Characteristics.Genes (Basel). 2022 Nov 27;13(12):2221. doi: 10.3390/genes13122221. Genes (Basel). 2022. PMID: 36553488 Free PMC article. Review.
-
Comparative genomics identifies candidate genes for infectious salmon anemia (ISA) resistance in Atlantic salmon (Salmo salar).Mar Biotechnol (NY). 2011 Apr;13(2):232-41. doi: 10.1007/s10126-010-9284-0. Epub 2010 Apr 16. Mar Biotechnol (NY). 2011. PMID: 20396924 Free PMC article.
-
Identification of a piscine reovirus-related pathogen in proliferative darkening syndrome (PDS) infected brown trout (Salmo trutta fario) using a next-generation technology detection pipeline.PLoS One. 2018 Oct 22;13(10):e0206164. doi: 10.1371/journal.pone.0206164. eCollection 2018. PLoS One. 2018. PMID: 30346982 Free PMC article.
-
The sockeye salmon genome, transcriptome, and analyses identifying population defining regions of the genome.PLoS One. 2020 Oct 29;15(10):e0240935. doi: 10.1371/journal.pone.0240935. eCollection 2020. PLoS One. 2020. PMID: 33119641 Free PMC article.
References
-
- Thorgaard GH, Bailey GS, Williams D, Buhler DR, Kaattari SL, Ristow SS, Hansen JD, Winton JR, Bartholomew JL, Nagler JJ, Walsh PJ, Vijayan MM, Devlin RH, Hardy RW, Overturf KE, Young WP, Robison BD, Rexroad C, Palti Y. Status and opportunities for genomics research with rainbow trout. Comp Biochem Physiol B Biochem Mol Biol. 2002;133:609–646. doi: 10.1016/S1096-4959(02)00167-7. - DOI - PubMed
-
- Nelson J. Fishes of the world. New York, Wiley; 2006.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

