Human Rad54 protein stimulates human Mus81-Eme1 endonuclease
- PMID: 19017809
- PMCID: PMC2587595
- DOI: 10.1073/pnas.0807016105
Human Rad54 protein stimulates human Mus81-Eme1 endonuclease
Abstract
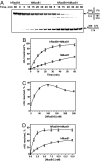
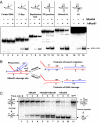
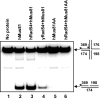
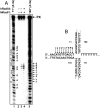
Rad54, a key protein of homologous recombination, physically interacts with a DNA structure-specific endonuclease, Mus81-Eme1. Genetic data indicate that Mus81-Eme1 and Rad54 might function together in the repair of damaged DNA. In vitro, Rad54 promotes branch migration of Holliday junctions, whereas the Mus81-Eme1 complex resolves DNA junctions by endonucleolytic cleavage. Here, we show that human Rad54 stimulates Mus81-Eme1 endonuclease activity on various Holliday junction-like intermediates. This stimulation is the product of specific interactions between the human Rad54 (hRad54) and Mus81 proteins, considering that Saccharomyces cerevisiae Rad54 protein does not stimulate human Mus81-Eme1 endonuclease activity. Stimulation of Mus81-Eme1 cleavage activity depends on formation of specific Rad54 complexes on DNA substrates occurring in the presence of ATP and, to a smaller extent, of other nucleotide cofactors. Thus, our results demonstrate a functional link between the branch migration activity of hRad54 and the structure-specific endonuclease activity of hMus81-Eme1, suggesting that the Rad54 and Mus81-Eme1 proteins may cooperate in the processing of Holliday junction-like intermediates during homologous recombination or DNA repair.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
The structure-specific endonuclease Mus81-Eme1 promotes conversion of interstrand DNA crosslinks into double-strands breaks.EMBO J. 2006 Oct 18;25(20):4921-32. doi: 10.1038/sj.emboj.7601344. Epub 2006 Oct 12. EMBO J. 2006. PMID: 17036055 Free PMC article.
-
Cooperativity of Mus81.Mms4 with Rad54 in the resolution of recombination and replication intermediates.J Biol Chem. 2009 Mar 20;284(12):7733-45. doi: 10.1074/jbc.M806192200. Epub 2009 Jan 7. J Biol Chem. 2009. Retraction in: J Biol Chem. 2024 Oct;300(10):107747. doi: 10.1016/j.jbc.2024.107747. PMID: 19129197 Free PMC article. Retracted.
-
Substrate specificity of the MUS81-EME2 structure selective endonuclease.Nucleic Acids Res. 2014 Apr;42(6):3833-45. doi: 10.1093/nar/gkt1333. Epub 2013 Dec 25. Nucleic Acids Res. 2014. PMID: 24371268 Free PMC article.
-
Exploring the roles of Mus81-Eme1/Mms4 at perturbed replication forks.DNA Repair (Amst). 2007 Jul 1;6(7):1004-17. doi: 10.1016/j.dnarep.2007.02.019. Epub 2007 Apr 3. DNA Repair (Amst). 2007. PMID: 17409028 Review.
-
The fuss about Mus81.Cell. 2001 Nov 30;107(5):551-4. doi: 10.1016/s0092-8674(01)00593-1. Cell. 2001. PMID: 11733053 Review.
Cited by
-
An Overview of the Molecular Mechanisms of Recombinational DNA Repair.Cold Spring Harb Perspect Biol. 2015 Nov 2;7(11):a016410. doi: 10.1101/cshperspect.a016410. Cold Spring Harb Perspect Biol. 2015. PMID: 26525148 Free PMC article. Review.
-
Junction resolving enzymes use multivalency to keep the Holliday junction dynamic.Nat Chem Biol. 2019 Mar;15(3):269-275. doi: 10.1038/s41589-018-0209-y. Epub 2019 Jan 21. Nat Chem Biol. 2019. PMID: 30664685 Free PMC article.
-
Esc2 promotes Mus81 complex-activity via its SUMO-like and DNA binding domains.Nucleic Acids Res. 2017 Jan 9;45(1):215-230. doi: 10.1093/nar/gkw882. Epub 2016 Sep 30. Nucleic Acids Res. 2017. PMID: 27694623 Free PMC article.
-
RAD54 promotes alternative lengthening of telomeres by mediating branch migration.EMBO Rep. 2020 Jun 4;21(6):e49495. doi: 10.15252/embr.201949495. Epub 2020 Apr 26. EMBO Rep. 2020. PMID: 32337843 Free PMC article.
-
MicroDNA levels are dependent on MMEJ, repressed by c-NHEJ pathway, and stimulated by DNA damage.Nucleic Acids Res. 2021 Nov 18;49(20):11787-11799. doi: 10.1093/nar/gkab984. Nucleic Acids Res. 2021. PMID: 34718766 Free PMC article.
References
-
- Sung P, Klein H. Mechanism of homologous recombination: Mediators and helicases take on regulatory functions. Nat Rev Mol Cell Biol. 2006;7:739–750. - PubMed
-
- Liu Y, West SC. Happy Hollidays: 40th anniversary of the Holliday junction. Nat Rev Mol Cell Biol. 2004;5:937–944. - PubMed
-
- Cox MM, et al. The importance of repairing stalled replication forks. Nature. 2000;404:37–41. - PubMed
-
- Seigneur M, Bidnenko V, Ehrlich SD, Michel B. RuvAB acts at arrested replication forks. Cell. 1998;95:419–430. - PubMed
-
- Lilley DM, White MF. The junction-resolving enzymes. Nat Rev Mol Cell Biol. 2001;2:433–443. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

