Architecture of thermal adaptation in an Exiguobacterium sibiricum strain isolated from 3 million year old permafrost: a genome and transcriptome approach
- PMID: 19019206
- PMCID: PMC2615787
- DOI: 10.1186/1471-2164-9-547
Architecture of thermal adaptation in an Exiguobacterium sibiricum strain isolated from 3 million year old permafrost: a genome and transcriptome approach
Abstract
Background: Many microorganisms have a wide temperature growth range and versatility to tolerate large thermal fluctuations in diverse environments, however not many have been fully explored over their entire growth temperature range through a holistic view of its physiology, genome, and transcriptome. We used Exiguobacterium sibiricum strain 255-15, a psychrotrophic bacterium from 3 million year old Siberian permafrost that grows from -5 degrees C to 39 degrees C to study its thermal adaptation.
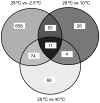
Results: The E. sibiricum genome has one chromosome and two small plasmids with a total of 3,015 protein-encoding genes (CDS), and a GC content of 47.7%. The genome and transcriptome analysis along with the organism's known physiology was used to better understand its thermal adaptation. A total of 27%, 3.2%, and 5.2% of E. sibiricum CDS spotted on the DNA microarray detected differentially expressed genes in cells grown at -2.5 degrees C, 10 degrees C, and 39 degrees C, respectively, when compared to cells grown at 28 degrees C. The hypothetical and unknown genes represented 10.6%, 0.89%, and 2.3% of the CDS differentially expressed when grown at -2.5 degrees C, 10 degrees C, and 39 degrees C versus 28 degrees C, respectively.
Conclusion: The results show that E. sibiricum is constitutively adapted to cold temperatures stressful to mesophiles since little differential gene expression was observed between 4 degrees C and 28 degrees C, but at the extremities of its Arrhenius growth profile, namely -2.5 degrees C and 39 degrees C, several physiological and metabolic adaptations associated with stress responses were observed.
Figures







Similar articles
-
Effect of low temperature and culture media on the growth and freeze-thawing tolerance of Exiguobacterium strains.Cryobiology. 2007 Apr;54(2):234-40. doi: 10.1016/j.cryobiol.2007.01.008. Epub 2007 Feb 6. Cryobiology. 2007. PMID: 17382311
-
Engineering of Thermal Stability in a Cold-Active Oligo-1,6-Glucosidase from Exiguobacterium sibiricum with Unusual Amino Acid Content.Biomolecules. 2021 Aug 17;11(8):1229. doi: 10.3390/biom11081229. Biomolecules. 2021. PMID: 34439895 Free PMC article.
-
Proteomic analysis of cold adaptation in a Siberian permafrost bacterium--Exiguobacterium sibiricum 255-15 by two-dimensional liquid separation coupled with mass spectrometry.Proteomics. 2006 Oct;6(19):5221-33. doi: 10.1002/pmic.200600071. Proteomics. 2006. PMID: 16955517
-
Microarray transcriptional profiling of Arctic Mesorhizobium strain N33 at low temperature provides insights into cold adaption strategies.BMC Genomics. 2015 May 15;16(1):383. doi: 10.1186/s12864-015-1611-4. BMC Genomics. 2015. PMID: 25975821 Free PMC article.
-
Conserved genomic and amino acid traits of cold adaptation in subzero-growing Arctic permafrost bacteria.FEMS Microbiol Ecol. 2018 Apr 1;94(4). doi: 10.1093/femsec/fiy023. FEMS Microbiol Ecol. 2018. PMID: 29528411
Cited by
-
Combined Methylome, Transcriptome and Proteome Analyses Document Rapid Acclimatization of a Bacterium to Environmental Changes.Front Microbiol. 2020 Sep 15;11:544785. doi: 10.3389/fmicb.2020.544785. eCollection 2020. Front Microbiol. 2020. PMID: 33042055 Free PMC article.
-
Draft Genome Sequence of Exiguobacterium sp. Strain BMC-KP, an Environmental Isolate from Bryn Mawr, Pennsylvania.Genome Announc. 2015 Oct 8;3(5):e01164-15. doi: 10.1128/genomeA.01164-15. Genome Announc. 2015. PMID: 26450734 Free PMC article.
-
Bacterial growth at -15 °C; molecular insights from the permafrost bacterium Planococcus halocryophilus Or1.ISME J. 2013 Jun;7(6):1211-26. doi: 10.1038/ismej.2013.8. Epub 2013 Feb 7. ISME J. 2013. PMID: 23389107 Free PMC article.
-
Survival and Energy Producing Strategies of Alkane Degraders Under Extreme Conditions and Their Biotechnological Potential.Front Microbiol. 2018 May 25;9:1081. doi: 10.3389/fmicb.2018.01081. eCollection 2018. Front Microbiol. 2018. PMID: 29910779 Free PMC article. Review.
-
Microbial Diversity in Extreme Marine Habitats and Their Biomolecules.Microorganisms. 2017 May 16;5(2):25. doi: 10.3390/microorganisms5020025. Microorganisms. 2017. PMID: 28509857 Free PMC article. Review.
References
-
- Gounot AM. Bacterial life at low temperature: physiological aspects and biotechnological implications. J Appl Bacteriol. 1991;71:386–397. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

